forked from AdamWilsonLabEDU/SpatialDataScience
-
Notifications
You must be signed in to change notification settings - Fork 0
Expand file tree
/
Copy pathPS_06_raster.Rmd
More file actions
187 lines (136 loc) · 3.27 KB
/
PS_06_raster.Rmd
File metadata and controls
187 lines (136 loc) · 3.27 KB
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
---
title: "Raster Analyses"
---
```{r echo=F, eval=T, message=F,warning=F}
library(sf)
library(spData)
library(viridis)
library(tidyverse)
library(raster)
```
---
## Available Spatial Packages
* `sp` First major spatial data package/format
* `rgdal` reading and writing spatial data
* `rgeos` Interface to open-source geometry engine (GEOS)
* `sf` Spatial Features in the 'tidyverse'
* `raster` gridded data (like satellite imagery)
* and a few others...
## Common Simple Feature (SF) types
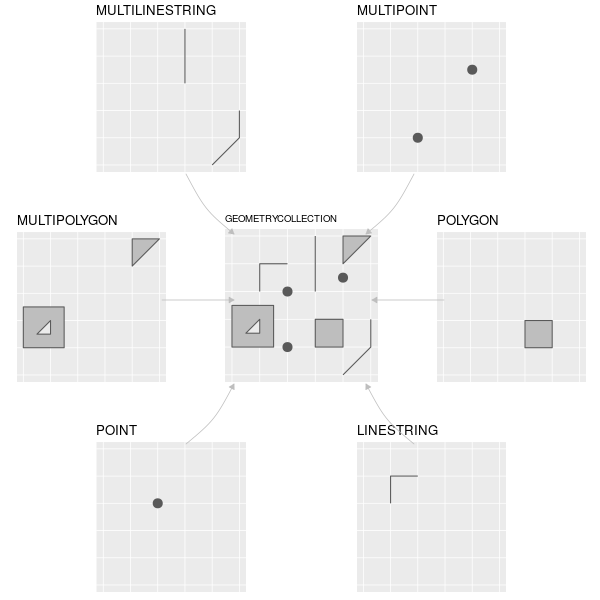
## What about grids?
```{r echo=F, message=F,warning=F}
library(raster)
library(spData)
data(elev) #load fake data from spData package
plot(elev)
```
# Raster Data
## Raster Data Types
Many types including:
* Thematic (classified/thematic) data: land-use or soils data.
* Continuous data (temperature, elevation, spectral)
* Imagery / Pictures (scanned maps, drawings, etc.)
Most Common:
* Satellite Imagery and derived products
* Earth System Models (e.g. climate models)
* Other models
## Raster data in the tidyverse
Raster data is not yet closely connected to the **tidyverse**, however:
- Some functions from `raster` work well in `pipes`
- Convert vector data to `Spatial*` form using `as(my_vector, "Spatial")` for raster-vector interactions
- Some early efforts to bring raster data into the **tidyverse**, including [tabularaster](https://github.com/hypertidy/tabularaster), [sfraster](https://github.com/mdsumner/sfraster), [fasterize](https://github.com/ecohealthalliance/fasterize), and [stars](https://github.com/r-spatial/stars) (multidimensional, large datasets).
## Raster Package
```{r message=F,warning=F}
library(raster)
library(spData)
data(elev) #load fake data from spData package
plot(elev) #load fake data from spData package
```
## Raster Data Structure
```{r}
elev
```
---
```{r}
str(elev)
```
---
### Arrays with metadata
```{r}
as.array(elev)
```
Rasters are just arrays / matricies with metadata.
---
### Cell Stats
```{r}
cellStats(elev, stat = mean, na.rm=T)
cellStats(elev, stat = quantile, na.rm=T)
```
---
### Raster Data Memory
```{r}
inMemory(elev)
canProcessInMemory(elev)
```
Raster does not try to load or work with large datasets in RAM. It creates temporary files and processes them in the background.
## Map Algebra
Convert feet to meters
```{r}
elev_m <- elev*0.3048
plot(elev_m)
```
---
### Simple Filter
```{r}
high_ground <- elev_m>8
plot(high_ground)
```
---
### Focal
```{r}
m <- matrix(1,nrow=3,ncol=3)
m
elev_smooth <- focal(elev_m,m,mean)
```
---
```{r}
par(mfrow=c(1,2))
plot(elev_m)
plot(elev_smooth)
```
## Arbitrary functions
```{r}
NA16=function(x) ifelse(x == 16,1,NA)
one_cell <- calc(elev,fun = NA16)
plot(one_cell)
```
## Distances to non-NA cells
```{r}
distance(one_cell)%>%
plot()
```
Distance unit is meters if RasterLayer is `+proj=longlat`, map units (typically also meters) otherwise.
## And much more
* aggregate (to coarser resolution)
* area (calculate cell area)
* buffer
* contour
* crop
* interpolate
* Moran's I
# Visualization
## ggplot
```{r, message=F}
library(rasterVis)
gplot(elev)+
geom_raster(aes(fill=value))
```
---
### All ggplot functionality available
```{r}
gplot(elev)+
geom_raster(aes(fill=value))+
coord_equal()+
scale_fill_viridis_c()
```