This section of the PyGAD's library documentation discusses the
pygad module.
Using the pygad module, instances of the genetic algorithm can be
created, run, saved, and loaded.
The first module available in PyGAD is named pygad and contains a
class named GA for building the genetic algorithm. The constructor,
methods, function, and attributes within the class are discussed in this
section.
For creating an instance of the pygad.GA class, the constructor
accepts several parameters that allow the user to customize the genetic
algorithm to different types of applications.
The pygad.GA class constructor supports the following parameters:
num_generations: Number of generations.num_parents_mating: Number of solutions to be selected as parents.fitness_func: Accepts a function/method and returns the fitness value of the solution. If a function is passed, then it must accept 3 parameters (1. the instance of thepygad.GAclass, 2. a single solution, and 3. its index in the population). If method, then it accepts a fourth parameter representing the method's class instance. Check the Preparing the fitness_func Parameter section for information about creating such a function.fitness_batch_size=None: A new optional parameter calledfitness_batch_sizeis supported to calculate the fitness function in batches. If it is assigned the value1orNone(default), then the normal flow is used where the fitness function is called for each individual solution. If thefitness_batch_sizeparameter is assigned a value satisfying this condition1 < fitness_batch_size <= sol_per_pop, then the solutions are grouped into batches of sizefitness_batch_sizeand the fitness function is called once for each batch. Check the Batch Fitness Calculation section for more details and examples. Added in from PyGAD 2.19.0.initial_population: A user-defined initial population. It is useful when the user wants to start the generations with a custom initial population. It defaults toNonewhich means no initial population is specified by the user. In this case, PyGAD creates an initial population using thesol_per_popandnum_genesparameters. An exception is raised if theinitial_populationisNonewhile any of the 2 parameters (sol_per_popornum_genes) is alsoNone. Introduced in PyGAD 2.0.0 and higher.sol_per_pop: Number of solutions (i.e. chromosomes) within the population. This parameter has no action ifinitial_populationparameter exists.num_genes: Number of genes in the solution/chromosome. This parameter is not needed if the user feeds the initial population to theinitial_populationparameter.gene_type=float: Controls the gene type. It can be assigned to a single data type that is applied to all genes or can specify the data type of each individual gene. It defaults tofloatwhich means all genes are offloatdata type. Starting from PyGAD 2.9.0, thegene_typeparameter can be assigned to a numeric value of any of these types:int,float, andnumpy.int/uint/float(8-64). Starting from PyGAD 2.14.0, it can be assigned to alist,tuple, or anumpy.ndarraywhich hold a data type for each gene (e.g.gene_type=[int, float, numpy.int8]). This helps to control the data type of each individual gene. In PyGAD 2.15.0, a precision for thefloatdata types can be specified (e.g.gene_type=[float, 2].init_range_low=-4: The lower value of the random range from which the gene values in the initial population are selected.init_range_lowdefaults to-4. Available in PyGAD 1.0.20 and higher. This parameter has no action if theinitial_populationparameter exists.init_range_high=4: The upper value of the random range from which the gene values in the initial population are selected.init_range_highdefaults to+4. Available in PyGAD 1.0.20 and higher. This parameter has no action if theinitial_populationparameter exists.parent_selection_type="sss": The parent selection type. Supported types aresss(for steady-state selection),rws(for roulette wheel selection),sus(for stochastic universal selection),rank(for rank selection),random(for random selection), andtournament(for tournament selection). A custom parent selection function can be passed starting from PyGAD 2.16.0. Check the User-Defined Crossover, Mutation, and Parent Selection Operators section for more details about building a user-defined parent selection function.keep_parents=-1: Number of parents to keep in the current population.-1(default) means to keep all parents in the next population.0means keep no parents in the next population. A valuegreater than 0means keeps the specified number of parents in the next population. Note that the value assigned tokeep_parentscannot be< - 1or greater than the number of solutions within the populationsol_per_pop. Starting from PyGAD 2.18.0, this parameter have an effect only when thekeep_elitismparameter is0. Starting from PyGAD 2.20.0, the parents' fitness from the last generation will not be re-used ifkeep_parents=0.keep_elitism=1: Added in PyGAD 2.18.0. It can take the value0or a positive integer that satisfies (0 <= keep_elitism <= sol_per_pop). It defaults to1which means only the best solution in the current generation is kept in the next generation. If assigned0, this means it has no effect. If assigned a positive integerK, then the bestKsolutions are kept in the next generation. It cannot be assigned a value greater than the value assigned to thesol_per_popparameter. If this parameter has a value different than0, then thekeep_parentsparameter will have no effect.K_tournament=3: In case that the parent selection type istournament, theK_tournamentspecifies the number of parents participating in the tournament selection. It defaults to3.crossover_type="single_point": Type of the crossover operation. Supported types aresingle_point(for single-point crossover),two_points(for two points crossover),uniform(for uniform crossover), andscattered(for scattered crossover). Scattered crossover is supported from PyGAD 2.9.0 and higher. It defaults tosingle_point. A custom crossover function can be passed starting from PyGAD 2.16.0. Check the User-Defined Crossover, Mutation, and Parent Selection Operators section for more details about creating a user-defined crossover function. Starting from PyGAD 2.2.2 and higher, ifcrossover_type=None, then the crossover step is bypassed which means no crossover is applied and thus no offspring will be created in the next generations. The next generation will use the solutions in the current population.crossover_probability=None: The probability of selecting a parent for applying the crossover operation. Its value must be between 0.0 and 1.0 inclusive. For each parent, a random value between 0.0 and 1.0 is generated. If this random value is less than or equal to the value assigned to thecrossover_probabilityparameter, then the parent is selected. Added in PyGAD 2.5.0 and higher.mutation_type="random": Type of the mutation operation. Supported types arerandom(for random mutation),swap(for swap mutation),inversion(for inversion mutation),scramble(for scramble mutation), andadaptive(for adaptive mutation). It defaults torandom. A custom mutation function can be passed starting from PyGAD 2.16.0. Check the User-Defined Crossover, Mutation, and Parent Selection Operators section for more details about creating a user-defined mutation function. Starting from PyGAD 2.2.2 and higher, ifmutation_type=None, then the mutation step is bypassed which means no mutation is applied and thus no changes are applied to the offspring created using the crossover operation. The offspring will be used unchanged in the next generation.Adaptivemutation is supported starting from PyGAD 2.10.0. For more information about adaptive mutation, go the the Adaptive Mutation section. For example about using adaptive mutation, check the Use Adaptive Mutation in PyGAD section.mutation_probability=None: The probability of selecting a gene for applying the mutation operation. Its value must be between 0.0 and 1.0 inclusive. For each gene in a solution, a random value between 0.0 and 1.0 is generated. If this random value is less than or equal to the value assigned to themutation_probabilityparameter, then the gene is selected. If this parameter exists, then there is no need for the 2 parametersmutation_percent_genesandmutation_num_genes. Added in PyGAD 2.5.0 and higher.mutation_by_replacement=False: An optional bool parameter. It works only when the selected type of mutation is random (mutation_type="random"). In this case,mutation_by_replacement=Truemeans replace the gene by the randomly generated value. If False, then it has no effect and random mutation works by adding the random value to the gene. Supported in PyGAD 2.2.2 and higher. Check the changes in PyGAD 2.2.2 under the Release History section for an example.mutation_percent_genes="default": Percentage of genes to mutate. It defaults to the string"default"which is later translated into the integer10which means 10% of the genes will be mutated. It must be>0and<=100. Out of this percentage, the number of genes to mutate is deduced which is assigned to themutation_num_genesparameter. Themutation_percent_genesparameter has no action ifmutation_probabilityormutation_num_genesexist. Starting from PyGAD 2.2.2 and higher, this parameter has no action ifmutation_typeisNone.mutation_num_genes=None: Number of genes to mutate which defaults toNonemeaning that no number is specified. Themutation_num_genesparameter has no action if the parametermutation_probabilityexists. Starting from PyGAD 2.2.2 and higher, this parameter has no action ifmutation_typeisNone.random_mutation_min_val=-1.0: Forrandommutation, therandom_mutation_min_valparameter specifies the start value of the range from which a random value is selected to be added to the gene. It defaults to-1. Starting from PyGAD 2.2.2 and higher, this parameter has no action ifmutation_typeisNone.random_mutation_max_val=1.0: Forrandommutation, therandom_mutation_max_valparameter specifies the end value of the range from which a random value is selected to be added to the gene. It defaults to+1. Starting from PyGAD 2.2.2 and higher, this parameter has no action ifmutation_typeisNone.gene_space=None: It is used to specify the possible values for each gene in case the user wants to restrict the gene values. It is useful if the gene space is restricted to a certain range or to discrete values. It accepts alist,range, ornumpy.ndarray. When all genes have the same global space, specify their values as alist/tuple/range/numpy.ndarray. For example,gene_space = [0.3, 5.2, -4, 8]restricts the gene values to the 4 specified values. If each gene has its own space, then thegene_spaceparameter can be nested like[[0.4, -5], [0.5, -3.2, 8.2, -9], ...]where the first sublist determines the values for the first gene, the second sublist for the second gene, and so on. If the nested list/tuple has aNonevalue, then the gene's initial value is selected randomly from the range specified by the 2 parametersinit_range_lowandinit_range_highand its mutation value is selected randomly from the range specified by the 2 parametersrandom_mutation_min_valandrandom_mutation_max_val.gene_spaceis added in PyGAD 2.5.0. Check the Release History of PyGAD 2.5.0 section of the documentation for more details. In PyGAD 2.9.0, NumPy arrays can be assigned to thegene_spaceparameter. In PyGAD 2.11.0, thegene_spaceparameter itself or any of its elements can be assigned to a dictionary to specify the lower and upper limits of the genes. For example,{'low': 2, 'high': 4}means the minimum and maximum values are 2 and 4, respectively. In PyGAD 2.15.0, a new key called"step"is supported to specify the step of moving from the start to the end of the range specified by the 2 existing keys"low"and"high".on_start=None: Accepts a function/method to be called only once before the genetic algorithm starts its evolution. If function, then it must accept a single parameter representing the instance of the genetic algorithm. If method, then it must accept 2 parameters where the second one refers to the method's object. Added in PyGAD 2.6.0.on_fitness=None: Accepts a function/method to be called after calculating the fitness values of all solutions in the population. If function, then it must accept 2 parameters: 1) a list of all solutions' fitness values 2) the instance of the genetic algorithm. If method, then it must accept 3 parameters where the third one refers to the method's object. Added in PyGAD 2.6.0.on_parents=None: Accepts a function/method to be called after selecting the parents that mates. If function, then it must accept 2 parameters: 1) the selected parents 2) the instance of the genetic algorithm If method, then it must accept 3 parameters where the third one refers to the method's object. Added in PyGAD 2.6.0.on_crossover=None: Accepts a function to be called each time the crossover operation is applied. This function must accept 2 parameters: the first one represents the instance of the genetic algorithm and the second one represents the offspring generated using crossover. Added in PyGAD 2.6.0.on_mutation=None: Accepts a function to be called each time the mutation operation is applied. This function must accept 2 parameters: the first one represents the instance of the genetic algorithm and the second one represents the offspring after applying the mutation. Added in PyGAD 2.6.0.on_generation=None: Accepts a function to be called after each generation. This function must accept a single parameter representing the instance of the genetic algorithm. If the function returned the stringstop, then therun()method stops without completing the other generations. Added in PyGAD 2.6.0.on_stop=None: Accepts a function to be called only once exactly before the genetic algorithm stops or when it completes all the generations. This function must accept 2 parameters: the first one represents the instance of the genetic algorithm and the second one is a list of fitness values of the last population's solutions. Added in PyGAD 2.6.0.delay_after_gen=0.0: It accepts a non-negative number specifying the time in seconds to wait after a generation completes and before going to the next generation. It defaults to0.0which means no delay after the generation. Available in PyGAD 2.4.0 and higher.save_best_solutions=False: WhenTrue, then the best solution after each generation is saved into an attribute namedbest_solutions. IfFalse(default), then no solutions are saved and thebest_solutionsattribute will be empty. Supported in PyGAD 2.9.0.save_solutions=False: IfTrue, then all solutions in each generation are appended into an attribute calledsolutionswhich is NumPy array. Supported in PyGAD 2.15.0.suppress_warnings=False: A bool parameter to control whether the warning messages are printed or not. It defaults toFalse.allow_duplicate_genes=True: Added in PyGAD 2.13.0. IfTrue, then a solution/chromosome may have duplicate gene values. IfFalse, then each gene will have a unique value in its solution.stop_criteria=None: Some criteria to stop the evolution. Added in PyGAD 2.15.0. Each criterion is passed asstrwhich has a stop word. The current 2 supported words arereachandsaturate.reachstops therun()method if the fitness value is equal to or greater than a given fitness value. An example forreachis"reach_40"which stops the evolution if the fitness is >= 40.saturatemeans stop the evolution if the fitness saturates for a given number of consecutive generations. An example forsaturateis"saturate_7"which means stop therun()method if the fitness does not change for 7 consecutive generations.parallel_processing=None: Added in PyGAD 2.17.0. IfNone(Default), this means no parallel processing is applied. It can accept a list/tuple of 2 elements [1) Can be either'process'or'thread'to indicate whether processes or threads are used, respectively., 2) The number of processes or threads to use.]. For example,parallel_processing=['process', 10]applies parallel processing with 10 processes. If a positive integer is assigned, then it is used as the number of threads. For example,parallel_processing=5uses 5 threads which is equivalent toparallel_processing=["thread", 5]. For more information, check the Parallel Processing in PyGAD section.random_seed=None: Added in PyGAD 2.18.0. It defines the random seed to be used by the random function generators (we use random functions in the NumPy and random modules). This helps to reproduce the same results by setting the same random seed (e.g.random_seed=2). If given the valueNone, then it has no effect.logger=None: Accepts an instance of thelogging.Loggerclass to log the outputs. Any message is no longer printed usingprint()but logged. Iflogger=None, then a logger is created that usesStreamHandlerto logs the messages to the console. Added in PyGAD 3.0.0. Check the Logging Outputs for more information.
The user doesn't have to specify all of such parameters while creating
an instance of the GA class. A very important parameter you must care
about is fitness_func which defines the fitness function.
It is OK to set the value of any of the 2 parameters init_range_low
and init_range_high to be equal, higher, or lower than the other
parameter (i.e. init_range_low is not needed to be lower than
init_range_high). The same holds for the random_mutation_min_val
and random_mutation_max_val parameters.
If the 2 parameters mutation_type and crossover_type are
None, this disables any type of evolution the genetic algorithm can
make. As a result, the genetic algorithm cannot find a better solution
that the best solution in the initial population.
The parameters are validated within the constructor. If at least a parameter is not correct, an exception is thrown.
plot_fitness(): Shows how the fitness evolves by generation.plot_genes(): Shows how the gene value changes for each generation.plot_new_solution_rate(): Shows the number of new solutions explored in each solution.
supported_int_types: A list of the supported types for the integer numbers.supported_float_types: A list of the supported types for the floating-point numbers.supported_int_float_types: A list of the supported types for all numbers. It just concatenates the previous 2 lists.
All the parameters and functions passed to the pygad.GA class
constructor are used as class attributes and methods in the instances of
the pygad.GA class. In addition to such attributes, there are other
attributes and methods added to the instances of the pygad.GA class:
The next 2 subsections list such attributes and methods.
generations_completed: Holds the number of the last completed generation.population: A NumPy array holding the initial population.valid_parameters: Set toTruewhen all the parameters passed in theGAclass constructor are valid.run_completed: Set toTrueonly after therun()method completes gracefully.pop_size: The population size.best_solutions_fitness: A list holding the fitness values of the best solutions for all generations.best_solution_generation: The generation number at which the best fitness value is reached. It is only assigned the generation number after therun()method completes. Otherwise, its value is -1.best_solutions: A NumPy array holding the best solution per each generation. It only exists when thesave_best_solutionsparameter in thepygad.GAclass constructor is set toTrue.last_generation_fitness: The fitness values of the solutions in the last generation. Added in PyGAD 2.12.0.previous_generation_fitness: At the end of each generation, the fitness of the most recent population is saved in thelast_generation_fitnessattribute. The fitness of the population exactly preceding this most recent population is saved in thelast_generation_fitnessattribute. Thisprevious_generation_fitnessattribute is used to fetch the pre-calculated fitness instead of calling the fitness function for already explored solutions. Added in PyGAD 2.16.2.last_generation_parents: The parents selected from the last generation. Added in PyGAD 2.12.0.last_generation_offspring_crossover: The offspring generated after applying the crossover in the last generation. Added in PyGAD 2.12.0.last_generation_offspring_mutation: The offspring generated after applying the mutation in the last generation. Added in PyGAD 2.12.0.gene_type_single: A flag that is set toTrueif thegene_typeparameter is assigned to a single data type that is applied to all genes. Ifgene_typeis assigned alist,tuple, ornumpy.ndarray, then the value ofgene_type_singlewill beFalse. Added in PyGAD 2.14.0.last_generation_parents_indices: This attribute holds the indices of the selected parents in the last generation. Supported in PyGAD 2.15.0.last_generation_elitism: This attribute holds the elitism of the last generation. It is effective only if thekeep_elitismparameter has a non-zero value. Supported in PyGAD 2.18.0.last_generation_elitism_indices: This attribute holds the indices of the elitism of the last generation. It is effective only if thekeep_elitismparameter has a non-zero value. Supported in PyGAD 2.19.0.logger: This attribute holds the logger from theloggingmodule. Supported in PyGAD 3.0.0.gene_space_unpacked: This is the unpacked version of thegene_spaceparameter. For example,range(1, 5)is unpacked to[1, 2, 3, 4]. For an infinite range like{'low': 2, 'high': 4}, then it is unpacked to a limited number of values (e.g. 100). Supported in PyGAD 3.1.0.
Note that the attributes with names starting with last_generation_
are updated after each generation.
cal_pop_fitness(): A method that calculates the fitness values for all solutions within the population by calling the function passed to thefitness_funcparameter for each solution.crossover(): Refers to the method that applies the crossover operator based on the selected type of crossover in thecrossover_typeproperty.mutation(): Refers to the method that applies the mutation operator based on the selected type of mutation in themutation_typeproperty.select_parents(): Refers to a method that selects the parents based on the parent selection type specified in theparent_selection_typeattribute.adaptive_mutation_population_fitness(): Returns the average fitness value used in the adaptive mutation to filter the solutions.solve_duplicate_genes_randomly(): Solves the duplicates in a solution by randomly selecting new values for the duplicating genes.solve_duplicate_genes_by_space(): Solves the duplicates in a solution by selecting values for the duplicating genes from the gene spaceunique_int_gene_from_range(): Finds a unique integer value for the gene.unique_genes_by_space(): Loops through all the duplicating genes to find unique values that from their gene spaces to solve the duplicates. For each duplicating gene, a call to theunique_gene_by_space()is made.unique_gene_by_space(): Returns a unique gene value for a single gene based on its value space to solve the duplicates.summary(): Prints a Keras-like summary of the PyGAD lifecycle. This helps to have an overview of the architecture. Supported in PyGAD 2.19.0. Check the Print Lifecycle Summary section for more details and examples.
The next sections discuss the methods available in the pygad.GA
class.
It creates an initial population randomly as a NumPy array. The array is
saved in the instance attribute named population.
Accepts the following parameters:
low: The lower value of the random range from which the gene values in the initial population are selected. It defaults to -4. Available in PyGAD 1.0.20 and higher.high: The upper value of the random range from which the gene values in the initial population are selected. It defaults to -4. Available in PyGAD 1.0.20.
This method assigns the values of the following 3 instance attributes:
pop_size: Size of the population.population: Initially, it holds the initial population and later updated after each generation.initial_population: Keeping the initial population.
The cal_pop_fitness() method calculates and returns the fitness
values of the solutions in the current population.
This function is optimized to save time by making fewer calls the fitness function. It follows this process:
- If the
save_solutionsparameter is set toTrue, then it checks if the solution is already explored and saved in thesolutionsinstance attribute. If so, then it just retrieves its fitness from thesolutions_fitnessinstance attribute without calling the fitness function. - If
save_solutionsis set toFalseor if it isTruebut the solution was not explored yet, then thecal_pop_fitness()method checks if thekeep_elitismparameter is set to a positive integer. If so, then it checks if the solution is saved into thelast_generation_elitisminstance attribute. If so, then it retrieves its fitness from theprevious_generation_fitnessinstance attribute. - If neither of the above 3 conditions apply (1.
save_solutionsis set toFalseor 2. if it isTruebut the solution was not explored yet or 3.keep_elitismis set to zero), then thecal_pop_fitness()method checks if thekeep_parentsparameter is set to-1or a positive integer. If so, then it checks if the solution is saved into thelast_generation_parentsinstance attribute. If so, then it retrieves its fitness from theprevious_generation_fitnessinstance attribute. - If neither of the above 4 conditions apply, then we have to call the
fitness function to calculate the fitness for the solution. This is
by calling the function assigned to the
fitness_funcparameter.
This function takes into consideration:
- The
parallel_processingparameter to check whether parallel processing is in effect. - The
fitness_batch_sizeparameter to check if the fitness should be calculated in batches of solutions.
It returns a vector of the solutions' fitness values.
Runs the genetic algorithm. This is the main method in which the genetic algorithm is evolved through some generations. It accepts no parameters as it uses the instance to access all of its requirements.
For each generation, the fitness values of all solutions within the
population are calculated according to the cal_pop_fitness() method
which internally just calls the function assigned to the
fitness_func parameter in the pygad.GA class constructor for
each solution.
According to the fitness values of all solutions, the parents are
selected using the select_parents() method. This method behaviour is
determined according to the parent selection type in the
parent_selection_type parameter in the pygad.GA class
constructor
Based on the selected parents, offspring are generated by applying the
crossover and mutation operations using the crossover() and
mutation() methods. The behaviour of such 2 methods is defined
according to the crossover_type and mutation_type parameters in
the pygad.GA class constructor.
After the generation completes, the following takes place:
- The
populationattribute is updated by the new population. - The
generations_completedattribute is assigned by the number of the last completed generation. - If there is a callback function assigned to the
on_generationattribute, then it will be called.
After the run() method completes, the following takes place:
- The
best_solution_generationis assigned the generation number at which the best fitness value is reached. - The
run_completedattribute is set toTrue.
The ParentSelection class in the pygad.utils.parent_selection
module has several methods for selecting the parents that will mate to
produce the offspring. All of such methods accept the same parameters
which are:
fitness: The fitness values of the solutions in the current population.num_parents: The number of parents to be selected.
All of such methods return an array of the selected parents.
The next subsections list the supported methods for parent selection.
Selects the parents using the steady-state selection technique.
Selects the parents using the rank selection technique.
Selects the parents randomly.
Selects the parents using the tournament selection technique.
Selects the parents using the roulette wheel selection technique.
Selects the parents using the stochastic universal selection technique.
The Crossover class in the pygad.utils.crossover module supports
several methods for applying crossover between the selected parents. All
of these methods accept the same parameters which are:
parents: The parents to mate for producing the offspring.offspring_size: The size of the offspring to produce.
All of such methods return an array of the produced offspring.
The next subsections list the supported methods for crossover.
Applies the single-point crossover. It selects a point randomly at which crossover takes place between the pairs of parents.
Applies the 2 points crossover. It selects the 2 points randomly at which crossover takes place between the pairs of parents.
Applies the uniform crossover. For each gene, a parent out of the 2 mating parents is selected randomly and the gene is copied from it.
Applies the scattered crossover. It randomly selects the gene from one of the 2 parents.
The Mutation class in the pygad.utils.mutation module supports
several methods for applying mutation. All of these methods accept the
same parameter which is:
offspring: The offspring to mutate.
All of such methods return an array of the mutated offspring.
The next subsections list the supported methods for mutation.
Applies the random mutation which changes the values of some genes
randomly. The number of genes is specified according to either the
mutation_num_genes or the mutation_percent_genes attributes.
For each gene, a random value is selected according to the range
specified by the 2 attributes random_mutation_min_val and
random_mutation_max_val. The random value is added to the selected
gene.
Applies the swap mutation which interchanges the values of 2 randomly selected genes.
Applies the inversion mutation which selects a subset of genes and inverts them.
Applies the scramble mutation which selects a subset of genes and shuffles their order randomly.
Applies the adaptive mutation which selects a subset of genes and shuffles their order randomly.
Returns information about the best solution found by the genetic algorithm.
It accepts the following parameters:
pop_fitness=None: An optional parameter that accepts a list of the fitness values of the solutions in the population. IfNone, then thecal_pop_fitness()method is called to calculate the fitness values of theself.population. Usega_instance.last_generation_fitnessto use latest fitness value and skip recalculation of the population fitness.
It returns the following:
best_solution: Best solution in the current population.best_solution_fitness: Fitness value of the best solution.best_match_idx: Index of the best solution in the current population.
Previously named plot_result(), this method creates, shows, and
returns a figure that summarizes how the fitness value evolves by
generation. It works only after completing at least 1 generation.
If no generation is completed (at least 1), an exception is raised.
Starting from PyGAD 2.15.0 and higher, this method accepts the following parameters:
title: Title of the figure.xlabel: X-axis label.ylabel: Y-axis label.linewidth: Line width of the plot. Defaults to3.font_size: Font size for the labels and title. Defaults to14.plot_type: Type of the plot which can be either"plot"(default),"scatter", or"bar".color: Color of the plot which defaults to"#3870FF".save_dir: Directory to save the figure.
The plot_new_solution_rate() method creates, shows, and returns a
figure that shows the number of new solutions explored in each
generation. This method works only when save_solutions=True in the
constructor of the pygad.GA class. It also works only after
completing at least 1 generation.
If no generation is completed (at least 1), an exception is raised.
This method accepts the following parameters:
title: Title of the figure.xlabel: X-axis label.ylabel: Y-axis label.linewidth: Line width of the plot. Defaults to3.font_size: Font size for the labels and title. Defaults to14.plot_type: Type of the plot which can be either"plot"(default),"scatter", or"bar".color: Color of the plot which defaults to"#3870FF".save_dir: Directory to save the figure.
The plot_genes() method creates, shows, and returns a figure that
describes each gene. It has different options to create the figures
which helps to:
- Explore the gene value for each generation by creating a normal plot.
- Create a histogram for each gene.
- Create a boxplot.
This is controlled by the graph_type parameter.
It works only after completing at least 1 generation. If no generation is completed, an exception is raised. If no generation is completed (at least 1), an exception is raised.
This method accepts the following parameters:
title: Title of the figure.xlabel: X-axis label.ylabel: Y-axis label.linewidth: Line width of the plot. Defaults to3.font_size: Font size for the labels and title. Defaults to14.plot_type: Type of the plot which can be either"plot"(default),"scatter", or"bar".graph_type: Type of the graph which can be either"plot"(default),"boxplot", or"histogram".fill_color: Fill color of the graph which defaults to"#3870FF". This has no effect ifgraph_type="plot".color: Color of the plot which defaults to"#3870FF".solutions: Defaults to"all"which means use all solutions. If"best"then only the best solutions are used.save_dir: Directory to save the figure.
An exception is raised if:
solutions="all"whilesave_solutions=Falsein the constructor of thepygad.GAclass. .solutions="best"whilesave_best_solutions=Falsein the constructor of thepygad.GAclass. .
Saves the genetic algorithm instance
Accepts the following parameter:
filename: Name of the file to save the instance. No extension is needed.
Besides the methods available in the pygad.GA class, this section
discusses the functions available in pygad. Up to this time, there
is only a single function named load().
Reads a saved instance of the genetic algorithm. This is not a method
but a function that is indented under the pygad module. So, it could
be called by the pygad module as follows: pygad.load(filename).
Accepts the following parameter:
filename: Name of the file holding the saved instance of the genetic algorithm. No extension is needed.
Returns the genetic algorithm instance.
To use the pygad module, here is a summary of the required steps:
- Preparing the
fitness_funcparameter. - Preparing Other Parameters.
- Import
pygad. - Create an Instance of the
pygad.GAClass. - Run the Genetic Algorithm.
- Plotting Results.
- Information about the Best Solution.
- Saving & Loading the Results.
Let's discuss how to do each of these steps.
Even there are some steps in the genetic algorithm pipeline that can work the same regardless of the problem being solved, one critical step is the calculation of the fitness value. There is no unique way of calculating the fitness value and it changes from one problem to another.
PyGAD has a parameter called fitness_func that allows the user to
specify a custom function/method to use when calculating the fitness.
This function/method must be a maximization function/method so that a
solution with a high fitness value returned is selected compared to a
solution with a low value. Doing that allows the user to freely use
PyGAD to solve any problem by passing the appropriate fitness
function/method. It is very important to understand the problem well for
creating it.
Let's discuss an example:
Given the following function:y = f(w1:w6) = w1x1 + w2x2 + w3x3 + w4x4 + w5x5 + 6wx6where (x1,x2,x3,x4,x5,x6)=(4, -2, 3.5, 5, -11, -4.7) and y=44What are the best values for the 6 weights (w1 to w6)? We are going to use the genetic algorithm to optimize this function.
So, the task is about using the genetic algorithm to find the best
values for the 6 weight W1 to W6. Thinking of the problem, it is
clear that the best solution is that returning an output that is close
to the desired output y=44. So, the fitness function/method should
return a value that gets higher when the solution's output is closer to
y=44. Here is a function that does that:
function_inputs = [4, -2, 3.5, 5, -11, -4.7] # Function inputs.
desired_output = 44 # Function output.
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / numpy.abs(output - desired_output)
return fitnessSuch a user-defined function must accept 3 parameters:
- The instance of the
pygad.GAclass. This helps the user to fetch any property that helps when calculating the fitness. - The solution(s) to calculate the fitness value(s). Note that the
fitness function can accept multiple solutions only if the
fitness_batch_sizeis given a value greater than 1. - The indices of the solutions in the population. The number of indices
also depends on the
fitness_batch_sizeparameter.
If a method is passed to the fitness_func parameter, then it accepts
a fourth parameter representing the method's instance.
The __code__ object is used to check if this function accepts the
required number of parameters. If more or fewer parameters are passed,
an exception is thrown.
By creating this function, you did a very important step towards using PyGAD.
Here is an example for preparing the other parameters:
num_generations = 50
num_parents_mating = 4
fitness_function = fitness_func
sol_per_pop = 8
num_genes = len(function_inputs)
init_range_low = -2
init_range_high = 5
parent_selection_type = "sss"
keep_parents = 1
crossover_type = "single_point"
mutation_type = "random"
mutation_percent_genes = 10An optional parameter named on_generation is supported which allows
the user to call a function (with a single parameter) after each
generation. Here is a simple function that just prints the current
generation number and the fitness value of the best solution in the
current generation. The generations_completed attribute of the GA
class returns the number of the last completed generation.
def on_gen(ga_instance):
print("Generation : ", ga_instance.generations_completed)
print("Fitness of the best solution :", ga_instance.best_solution()[1])After being defined, the function is assigned to the on_generation
parameter of the GA class constructor. By doing that, the on_gen()
function will be called after each generation.
ga_instance = pygad.GA(...,
on_generation=on_gen,
...)After the parameters are prepared, we can import PyGAD and build an
instance of the pygad.GA class.
The next step is to import PyGAD as follows:
import pygadThe pygad.GA class holds the implementation of all methods for
running the genetic algorithm.
The pygad.GA class is instantiated where the previously prepared
parameters are fed to its constructor. The constructor is responsible
for creating the initial population.
ga_instance = pygad.GA(num_generations=num_generations,
num_parents_mating=num_parents_mating,
fitness_func=fitness_function,
sol_per_pop=sol_per_pop,
num_genes=num_genes,
init_range_low=init_range_low,
init_range_high=init_range_high,
parent_selection_type=parent_selection_type,
keep_parents=keep_parents,
crossover_type=crossover_type,
mutation_type=mutation_type,
mutation_percent_genes=mutation_percent_genes)After an instance of the pygad.GA class is created, the next step is
to call the run() method as follows:
ga_instance.run()Inside this method, the genetic algorithm evolves over some generations by doing the following tasks:
- Calculating the fitness values of the solutions within the current population.
- Select the best solutions as parents in the mating pool.
- Apply the crossover & mutation operation
- Repeat the process for the specified number of generations.
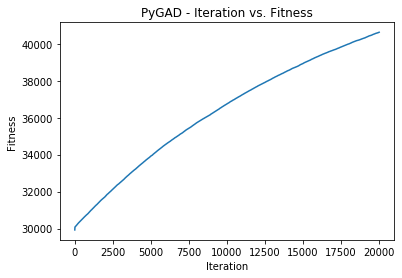
There is a method named plot_fitness() which creates a figure
summarizing how the fitness values of the solutions change with the
generations.
ga_instance.plot_fitness()The following information about the best solution in the last population
is returned using the best_solution() method.
- Solution
- Fitness value of the solution
- Index of the solution within the population
solution, solution_fitness, solution_idx = ga_instance.best_solution()
print("Parameters of the best solution : {solution}".format(solution=solution))
print("Fitness value of the best solution = {solution_fitness}".format(solution_fitness=solution_fitness))
print("Index of the best solution : {solution_idx}".format(solution_idx=solution_idx))Using the best_solution_generation attribute of the instance from
the pygad.GA class, the generation number at which the
best fitness is reached could be fetched.
if ga_instance.best_solution_generation != -1:
print("Best fitness value reached after {best_solution_generation} generations.".format(best_solution_generation=ga_instance.best_solution_generation))After the run() method completes, it is possible to save the current
instance of the genetic algorithm to avoid losing the progress made. The
save() method is available for that purpose. Just pass the file name
to it without an extension. According to the next code, a file named
genetic.pkl will be created and saved in the current directory.
filename = 'genetic'
ga_instance.save(filename=filename)You can also load the saved model using the load() function and
continue using it. For example, you might run the genetic algorithm for
some generations, save its current state using the save() method,
load the model using the load() function, and then call the
run() method again.
loaded_ga_instance = pygad.load(filename=filename)After the instance is loaded, you can use it to run any method or access any property.
print(loaded_ga_instance.best_solution())PyGAD supports different types for selecting the parents and applying
the crossover & mutation operators. More features will be added in the
future. To ask for a new feature, please check the Ask for Feature
section.
The supported crossover operations at this time are:
- Single point: Implemented using the
single_point_crossover()method. - Two points: Implemented using the
two_points_crossover()method. - Uniform: Implemented using the
uniform_crossover()method.
The supported mutation operations at this time are:
- Random: Implemented using the
random_mutation()method. - Swap: Implemented using the
swap_mutation()method. - Inversion: Implemented using the
inversion_mutation()method. - Scramble: Implemented using the
scramble_mutation()method.
The supported parent selection techniques at this time are:
- Steady-state: Implemented using the
steady_state_selection()method. - Roulette wheel: Implemented using the
roulette_wheel_selection()method. - Stochastic universal: Implemented using the
stochastic_universal_selection()method. - Rank: Implemented using the
rank_selection()method. - Random: Implemented using the
random_selection()method. - Tournament: Implemented using the
tournament_selection()method.
The next figure lists the different stages in the lifecycle of an
instance of the pygad.GA class. Note that PyGAD stops when either
all generations are completed or when the function passed to the
on_generation parameter returns the string stop.
The next code implements all the callback functions to trace the execution of the genetic algorithm. Each callback function prints its name.
import pygad
import numpy
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
fitness_function = fitness_func
def on_start(ga_instance):
print("on_start()")
def on_fitness(ga_instance, population_fitness):
print("on_fitness()")
def on_parents(ga_instance, selected_parents):
print("on_parents()")
def on_crossover(ga_instance, offspring_crossover):
print("on_crossover()")
def on_mutation(ga_instance, offspring_mutation):
print("on_mutation()")
def on_generation(ga_instance):
print("on_generation()")
def on_stop(ga_instance, last_population_fitness):
print("on_stop()")
ga_instance = pygad.GA(num_generations=3,
num_parents_mating=5,
fitness_func=fitness_function,
sol_per_pop=10,
num_genes=len(function_inputs),
on_start=on_start,
on_fitness=on_fitness,
on_parents=on_parents,
on_crossover=on_crossover,
on_mutation=on_mutation,
on_generation=on_generation,
on_stop=on_stop)
ga_instance.run()Based on the used 3 generations as assigned to the num_generations
argument, here is the output.
on_start() on_fitness() on_parents() on_crossover() on_mutation() on_generation() on_fitness() on_parents() on_crossover() on_mutation() on_generation() on_fitness() on_parents() on_crossover() on_mutation() on_generation() on_stop()
In the regular genetic algorithm, the mutation works by selecting a single fixed mutation rate for all solutions regardless of their fitness values. So, regardless on whether this solution has high or low quality, the same number of genes are mutated all the time.
The pitfalls of using a constant mutation rate for all solutions are summarized in this paper Libelli, S. Marsili, and P. Alba. "Adaptive mutation in genetic algorithms." Soft computing 4.2 (2000): 76-80 as follows:
The weak point of "classical" GAs is the total randomness of mutation, which is applied equally to all chromosomes, irrespective of their fitness. Thus a very good chromosome is equally likely to be disrupted by mutation as a bad one.
On the other hand, bad chromosomes are less likely to produce good ones through crossover, because of their lack of building blocks, until they remain unchanged. They would benefit the most from mutation and could be used to spread throughout the parameter space to increase the search thoroughness. So there are two conflicting needs in determining the best probability of mutation.
Usually, a reasonable compromise in the case of a constant mutation is to keep the probability low to avoid disruption of good chromosomes, but this would prevent a high mutation rate of low-fitness chromosomes. Thus a constant probability of mutation would probably miss both goals and result in a slow improvement of the population.
According to Libelli, S. Marsili, and P. Alba. work, the adaptive mutation solves the problems of constant mutation.
Adaptive mutation works as follows:
- Calculate the average fitness value of the population (
f_avg). - For each chromosome, calculate its fitness value (
f). - If
f<f_avg, then this solution is regarded as a low-quality solution and thus the mutation rate should be kept high because this would increase the quality of this solution. - If
f>f_avg, then this solution is regarded as a high-quality solution and thus the mutation rate should be kept low to avoid disrupting this high quality solution.
In PyGAD, if f=f_avg, then the solution is regarded of high quality.
The next figure summarizes the previous steps.
This strategy is applied in PyGAD.
In PyGAD 2.10.0, adaptive mutation is supported. To use it, just follow the following 2 simple steps:
- In the constructor of the
pygad.GAclass, setmutation_type="adaptive"to specify that the type of mutation is adaptive. - Specify the mutation rates for the low and high quality solutions
using one of these 3 parameters according to your preference:
mutation_probability,mutation_num_genes, andmutation_percent_genes. Please check the documentation of each of these parameters for more information.
When adaptive mutation is used, then the value assigned to any of the 3 parameters can be of any of these data types:
listtuplenumpy.ndarray
Whatever the data type used, the length of the list, tuple, or
the numpy.ndarray must be exactly 2. That is there are just 2
values:
- The first value is the mutation rate for the low-quality solutions.
- The second value is the mutation rate for the high-quality solutions.
PyGAD expects that the first value is higher than the second value and thus a warning is printed in case the first value is lower than the second one.
Here are some examples to feed the mutation rates:
# mutation_probability
mutation_probability = [0.25, 0.1]
mutation_probability = (0.35, 0.17)
mutation_probability = numpy.array([0.15, 0.05])
# mutation_num_genes
mutation_num_genes = [4, 2]
mutation_num_genes = (3, 1)
mutation_num_genes = numpy.array([7, 2])
# mutation_percent_genes
mutation_percent_genes = [25, 12]
mutation_percent_genes = (15, 8)
mutation_percent_genes = numpy.array([21, 13])Assume that the average fitness is 12 and the fitness values of 2 solutions are 15 and 7. If the mutation probabilities are specified as follows:
mutation_probability = [0.25, 0.1]Then the mutation probability of the first solution is 0.1 because its fitness is 15 which is higher than the average fitness 12. The mutation probability of the second solution is 0.25 because its fitness is 7 which is lower than the average fitness 12.
Here is an example that uses adaptive mutation.
import pygad
import numpy
function_inputs = [4,-2,3.5,5,-11,-4.7] # Function inputs.
desired_output = 44 # Function output.
def fitness_func(ga_instance, solution, solution_idx):
# The fitness function calulates the sum of products between each input and its corresponding weight.
output = numpy.sum(solution*function_inputs)
# The value 0.000001 is used to avoid the Inf value when the denominator numpy.abs(output - desired_output) is 0.0.
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
# Creating an instance of the GA class inside the ga module. Some parameters are initialized within the constructor.
ga_instance = pygad.GA(num_generations=200,
fitness_func=fitness_func,
num_parents_mating=10,
sol_per_pop=20,
num_genes=len(function_inputs),
mutation_type="adaptive",
mutation_num_genes=(3, 1))
# Running the GA to optimize the parameters of the function.
ga_instance.run()
ga_instance.plot_fitness(title="PyGAD with Adaptive Mutation", linewidth=5)In PyGAD
2.11.0,
the gene_space parameter supported a new feature to allow
customizing the range of accepted values for each gene. Let's take a
quick review of the gene_space parameter to build over it.
The gene_space parameter allows the user to feed the space of values
of each gene. This way the accepted values for each gene is retracted to
the user-defined values. Assume there is a problem that has 3 genes
where each gene has different set of values as follows:
- Gene 1:
[0.4, 12, -5, 21.2] - Gene 2:
[-2, 0.3] - Gene 3:
[1.2, 63.2, 7.4]
Then, the gene_space for this problem is as given below. Note that
the order is very important.
gene_space = [[0.4, 12, -5, 21.2],
[-2, 0.3],
[1.2, 63.2, 7.4]]In case all genes share the same set of values, then simply feed a
single list to the gene_space parameter as follows. In this case,
all genes can only take values from this list of 6 values.
gene_space = [33, 7, 0.5, 95. 6.3, 0.74]The previous example restricts the gene values to just a set of fixed
number of discrete values. In case you want to use a range of discrete
values to the gene, then you can use the range() function. For
example, range(1, 7) means the set of allowed values for the gene
are 1, 2, 3, 4, 5, and 6. You can also use the numpy.arange() or
numpy.linspace() functions for the same purpose.
The previous discussion only works with a range of discrete values not
continuous values. In PyGAD
2.11.0,
the gene_space parameter can be assigned a dictionary that allows
the gene to have values from a continuous range.
Assuming you want to restrict the gene within this half-open range [1 to 5) where 1 is included and 5 is not. Then simply create a dictionary with 2 items where the keys of the 2 items are:
'low': The minimum value in the range which is 1 in the example.'high': The maximum value in the range which is 5 in the example.
The dictionary will look like that:
{'low': 1,
'high': 5}It is not acceptable to add more than 2 items in the dictionary or use
other keys than 'low' and 'high'.
For a 3-gene problem, the next code creates a dictionary for each gene to restrict its values in a continuous range. For the first gene, it can take any floating-point value from the range that starts from 1 (inclusive) and ends at 5 (exclusive).
gene_space = [{'low': 1, 'high': 5}, {'low': 0.3, 'high': 1.4}, {'low': -0.2, 'high': 4.5}]The gene_space parameter customizes the space of values of each
gene.
Assuming that all genes have the same global space which include the
values 0.3, 5.2, -4, and 8, then those values can be assigned to the
gene_space parameter as a list, tuple, or range. Here is a list
assigned to this parameter. By doing that, then the gene values are
restricted to those assigned to the gene_space parameter.
gene_space = [0.3, 5.2, -4, 8]If some genes have different spaces, then gene_space should accept a
nested list or tuple. In this case, the elements could be:
- Number (of
int,float, orNumPydata types): A single value to be assigned to the gene. This means this gene will have the same value across all generations. list,tuple,numpy.ndarray, or any range likerange,numpy.arange(), ornumpy.linspace: It holds the space for each individual gene. But this space is usually discrete. That is there is a set of finite values to select from.dict: To sample a value for a gene from a continuous range. The dictionary must have 2 mandatory keys which are"low"and"high"in addition to an optional key which is"step". A random value is returned between the values assigned to the items with"low"and"high"keys. If the"step"exists, then this works as the previous options (i.e. discrete set of values).None: A gene with its space set toNoneis initialized randomly from the range specified by the 2 parametersinit_range_lowandinit_range_high. For mutation, its value is mutated based on a random value from the range specified by the 2 parametersrandom_mutation_min_valandrandom_mutation_max_val. If all elements in thegene_spaceparameter areNone, the parameter will not have any effect.
Assuming that a chromosome has 2 genes and each gene has a different
value space. Then the gene_space could be assigned a nested
list/tuple where each element determines the space of a gene.
According to the next code, the space of the first gene is [0.4, -5]
which has 2 values and the space for the second gene is
[0.5, -3.2, 8.8, -9] which has 4 values.
gene_space = [[0.4, -5], [0.5, -3.2, 8.2, -9]]For a 2 gene chromosome, if the first gene space is restricted to the discrete values from 0 to 4 and the second gene is restricted to the values from 10 to 19, then it could be specified according to the next code.
gene_space = [range(5), range(10, 20)]The gene_space can also be assigned to a single range, as given
below, where the values of all genes are sampled from the same range.
gene_space = numpy.arange(15)The gene_space can be assigned a dictionary to sample a value from a
continuous range.
gene_space = {"low": 4, "high": 30}A step also can be assigned to the dictionary. This works as if a range is used.
gene_space = {"low": 4, "high": 30, "step": 2.5}Setting adictlike{"low": 0, "high": 10}in thegene_spacemeans that random values from the continuous range [0, 10) are sampled. Note that0is included but10is not included while sampling. Thus, the maximum value that could be returned is less than10like9.9999. But if the user decided to round the genes using, for example,[float, 2], then this value will become 10. So, the user should be careful to the inputs.
If a None is assigned to only a single gene, then its value will be
randomly generated initially using the init_range_low and
init_range_high parameters in the pygad.GA class's constructor.
During mutation, the value are sampled from the range defined by the 2
parameters random_mutation_min_val and random_mutation_max_val.
This is an example where the second gene is given a None value.
gene_space = [range(5), None, numpy.linspace(10, 20, 300)]If the user did not assign the initial population to the
initial_population parameter, the initial population is created
randomly based on the gene_space parameter. Moreover, the mutation
is applied based on this parameter.
If a gene has its static space defined in the gene_space parameter,
then mutation works by replacing the gene value by a value randomly
selected from the gene space. This happens for both int and
float data types.
For example, the following gene_space has the static space
[1, 2, 3] defined for the first gene. So, this gene can only have a
value out of these 3 values.
Gene space: [[1, 2, 3],
None]
Solution: [1, 5]For a solution like [1, -0.5, 4], then mutation happens for the
first gene by simply replacing its current value by a randomly selected
value (other than its current value if possible). So, the value 1 will
be replaced by either 2 or 3.
For the second gene, its space is set to None. So, traditional
mutation happens for this gene by:
- Generating a random value from the range defined by the
random_mutation_min_valandrandom_mutation_max_valparameters. - Adding this random value to the current gene's value.
If its current value is 5 and the random value is -0.5, then the new
value is 4.5. If the gene type is integer, then the value will be
rounded.
In PyGAD
2.4.0,
it is possible to stop the genetic algorithm after any generation. All
you need to do it to return the string "stop" in the callback
function on_generation. When this callback function is implemented
and assigned to the on_generation parameter in the constructor of
the pygad.GA class, then the algorithm immediately stops after
completing its current generation. Let's discuss an example.
Assume that the user wants to stop algorithm either after the 100
generations or if a condition is met. The user may assign a value of 100
to the num_generations parameter of the pygad.GA class
constructor.
The condition that stops the algorithm is written in a callback function
like the one in the next code. If the fitness value of the best solution
exceeds 70, then the string "stop" is returned.
def func_generation(ga_instance):
if ga_instance.best_solution()[1] >= 70:
return "stop"In PyGAD
2.15.0,
a new parameter named stop_criteria is added to the constructor of
the pygad.GA class. It helps to stop the evolution based on some
criteria. It can be assigned to one or more criterion.
Each criterion is passed as str that consists of 2 parts:
- Stop word.
- Number.
It takes this form:
"word_num"The current 2 supported words are reach and saturate.
The reach word stops the run() method if the fitness value is
equal to or greater than a given fitness value. An example for reach
is "reach_40" which stops the evolution if the fitness is >= 40.
saturate stops the evolution if the fitness saturates for a given
number of consecutive generations. An example for saturate is
"saturate_7" which means stop the run() method if the fitness
does not change for 7 consecutive generations.
Here is an example that stops the evolution if either the fitness value
reached 127.4 or if the fitness saturates for 15 generations.
import pygad
import numpy
equation_inputs = [4, -2, 3.5, 8, 9, 4]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=200,
sol_per_pop=10,
num_parents_mating=4,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
stop_criteria=["reach_127.4", "saturate_15"])
ga_instance.run()
print("Number of generations passed is {generations_completed}".format(generations_completed=ga_instance.generations_completed))In PyGAD
2.18.0,
a new parameter called keep_elitism is supported. It accepts an
integer to define the number of elitism (i.e. best solutions) to keep in
the next generation. This parameter defaults to 1 which means only
the best solution is kept in the next generation.
In the next example, the keep_elitism parameter in the constructor
of the pygad.GA class is set to 2. Thus, the best 2 solutions in
each generation are kept in the next generation.
import numpy
import pygad
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / numpy.abs(output - desired_output)
return fitness
ga_instance = pygad.GA(num_generations=2,
num_parents_mating=3,
fitness_func=fitness_func,
num_genes=6,
sol_per_pop=5,
keep_elitism=2)
ga_instance.run()The value passed to the keep_elitism parameter must satisfy 2
conditions:
- It must be
>= 0. - It must be
<= sol_per_pop. That is its value cannot exceed the number of solutions in the current population.
In the previous example, if the keep_elitism parameter is set equal
to the value passed to the sol_per_pop parameter, which is 5, then
there will be no evolution at all as in the next figure. This is because
all the 5 solutions are used as elitism in the next generation and no
offspring will be created.
...
ga_instance = pygad.GA(...,
sol_per_pop=5,
keep_elitism=5)
ga_instance.run()Note that if the keep_elitism parameter is effective (i.e. is
assigned a positive integer, not zero), then the keep_parents
parameter will have no effect. Because the default value of the
keep_elitism parameter is 1, then the keep_parents parameter has
no effect by default. The keep_parents parameter is only effective
when keep_elitism=0.
In PyGAD
2.18.0,
a new parameter called random_seed is supported. Its value is used
as a seed for the random function generators.
PyGAD uses random functions in these 2 libraries:
- NumPy
- random
The random_seed parameter defaults to None which means no seed
is used. As a result, different random numbers are generated for each
run of PyGAD.
If this parameter is assigned a proper seed, then the results will be reproducible. In the next example, the integer 2 is used as a random seed.
import numpy
import pygad
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / numpy.abs(output - desired_output)
return fitness
ga_instance = pygad.GA(num_generations=2,
num_parents_mating=3,
fitness_func=fitness_func,
sol_per_pop=5,
num_genes=6,
random_seed=2)
ga_instance.run()
best_solution, best_solution_fitness, best_match_idx = ga_instance.best_solution()
print(best_solution)
print(best_solution_fitness)This is the best solution found and its fitness value.
[ 2.77249188 -4.06570662 0.04196872 -3.47770796 -0.57502138 -3.22775267] 0.04872203136549972
After running the code again, it will find the same result.
[ 2.77249188 -4.06570662 0.04196872 -3.47770796 -0.57502138 -3.22775267] 0.04872203136549972
In PyGAD
2.18.0,
and thanks for Felix Bernhard for
opening this GitHub
issue,
the values of these 4 instance attributes are no longer reset after each
call to the run() method.
self.best_solutionsself.best_solutions_fitnessself.solutionsself.solutions_fitness
This helps the user to continue where the last run stopped without loosing the values of these 4 attributes.
Now, the user can save the model by calling the save() method.
import pygad
def fitness_func(ga_instance, solution, solution_idx):
...
return fitness
ga_instance = pygad.GA(...)
ga_instance.run()
ga_instance.plot_fitness()
ga_instance.save("pygad_GA")Then the saved model is loaded by calling the load() function. After
calling the run() method over the loaded instance, then the data
from the previous 4 attributes are not reset but extended with the new
data.
import pygad
def fitness_func(ga_instance, solution, solution_idx):
...
return fitness
loaded_ga_instance = pygad.load("pygad_GA")
loaded_ga_instance.run()
loaded_ga_instance.plot_fitness()The plot created by the plot_fitness() method will show the data
collected from both the runs.
Note that the 2 attributes (self.best_solutions and
self.best_solutions_fitness) only work if the
save_best_solutions parameter is set to True. Also, the 2
attributes (self.solutions and self.solutions_fitness) only work
if the save_solutions parameter is True.
In PyGAD
2.13.0,
a new bool parameter called allow_duplicate_genes is supported to
control whether duplicates are supported in the chromosome or not. In
other words, whether 2 or more genes might have the same exact value.
If allow_duplicate_genes=True (which is the default case), genes may
have the same value. If allow_duplicate_genes=False, then no 2 genes
will have the same value given that there are enough unique values for
the genes.
The next code gives an example to use the allow_duplicate_genes
parameter. A callback generation function is implemented to print the
population after each generation.
import pygad
def fitness_func(ga_instance, solution, solution_idx):
return 0
def on_generation(ga):
print("Generation", ga.generations_completed)
print(ga.population)
ga_instance = pygad.GA(num_generations=5,
sol_per_pop=5,
num_genes=4,
mutation_num_genes=3,
random_mutation_min_val=-5,
random_mutation_max_val=5,
num_parents_mating=2,
fitness_func=fitness_func,
gene_type=int,
on_generation=on_generation,
allow_duplicate_genes=False)
ga_instance.run()Here are the population after the 5 generations. Note how there are no duplicate values.
Generation 1
[[ 2 -2 -3 3]
[ 0 1 2 3]
[ 5 -3 6 3]
[-3 1 -2 4]
[-1 0 -2 3]]
Generation 2
[[-1 0 -2 3]
[-3 1 -2 4]
[ 0 -3 -2 6]
[-3 0 -2 3]
[ 1 -4 2 4]]
Generation 3
[[ 1 -4 2 4]
[-3 0 -2 3]
[ 4 0 -2 1]
[-4 0 -2 -3]
[-4 2 0 3]]
Generation 4
[[-4 2 0 3]
[-4 0 -2 -3]
[-2 5 4 -3]
[-1 2 -4 4]
[-4 2 0 -3]]
Generation 5
[[-4 2 0 -3]
[-1 2 -4 4]
[ 3 4 -4 0]
[-1 0 2 -2]
[-4 2 -1 1]]The allow_duplicate_genes parameter is configured with use with the
gene_space parameter. Here is an example where each of the 4 genes
has the same space of values that consists of 4 values (1, 2, 3, and 4).
import pygad
def fitness_func(ga_instance, solution, solution_idx):
return 0
def on_generation(ga):
print("Generation", ga.generations_completed)
print(ga.population)
ga_instance = pygad.GA(num_generations=1,
sol_per_pop=5,
num_genes=4,
num_parents_mating=2,
fitness_func=fitness_func,
gene_type=int,
gene_space=[[1, 2, 3, 4], [1, 2, 3, 4], [1, 2, 3, 4], [1, 2, 3, 4]],
on_generation=on_generation,
allow_duplicate_genes=False)
ga_instance.run()Even that all the genes share the same space of values, no 2 genes duplicate their values as provided by the next output.
Generation 1
[[2 3 1 4]
[2 3 1 4]
[2 4 1 3]
[2 3 1 4]
[1 3 2 4]]
Generation 2
[[1 3 2 4]
[2 3 1 4]
[1 3 2 4]
[2 3 4 1]
[1 3 4 2]]
Generation 3
[[1 3 4 2]
[2 3 4 1]
[1 3 4 2]
[3 1 4 2]
[3 2 4 1]]
Generation 4
[[3 2 4 1]
[3 1 4 2]
[3 2 4 1]
[1 2 4 3]
[1 3 4 2]]
Generation 5
[[1 3 4 2]
[1 2 4 3]
[2 1 4 3]
[1 2 4 3]
[1 2 4 3]]You should care of giving enough values for the genes so that PyGAD is able to find alternatives for the gene value in case it duplicates with another gene.
There might be 2 duplicate genes where changing either of the 2
duplicating genes will not solve the problem. For example, if
gene_space=[[3, 0, 1], [4, 1, 2], [0, 2], [3, 2, 0]] and the
solution is [3 2 0 0], then the values of the last 2 genes
duplicate. There are no possible changes in the last 2 genes to solve
the problem.
This problem can be solved by randomly changing one of the
non-duplicating genes that may make a room for a unique value in one the
2 duplicating genes. For example, by changing the second gene from 2 to
4, then any of the last 2 genes can take the value 2 and solve the
duplicates. The resultant gene is then [3 4 2 0]. But this option is
not yet supported in PyGAD.
When allow_duplicate_genes=False and a user-defined gene_space
is used, it sometimes happen that there is no room to solve the
duplicates between the 2 genes by simply replacing the value of one gene
by another gene. In PyGAD
3.1.0,
the duplicates are solved by looking for a third gene that will help in
solving the duplicates. The following examples explain how it works.
Example 1:
Let's assume that this gene space is used and there is a solution with 2 duplicate genes with the same value 4.
Gene space: [[2, 3],
[3, 4],
[4, 5],
[5, 6]]
Solution: [3, 4, 4, 5]By checking the gene space, the second gene can have the values
[3, 4] and the third gene can have the values [4, 5]. To solve
the duplicates, we have the value of any of these 2 genes.
If the value of the second gene changes from 4 to 3, then it will be duplicate with the first gene. If we are to change the value of the third gene from 4 to 5, then it will duplicate with the fourth gene. As a conclusion, trying to just selecting a different gene value for either the second or third genes will introduce new duplicating genes.
When there are 2 duplicate genes but there is no way to solve their duplicates, then the solution is to change a third gene that makes a room to solve the duplicates between the 2 genes.
In our example, duplicates between the second and third genes can be solved by, for example,:
- Changing the first gene from 3 to 2 then changing the second gene from 4 to 3.
- Or changing the fourth gene from 5 to 6 then changing the third gene from 4 to 5.
Generally, this is how to solve such duplicates:
- For any duplicate gene GENE1, select another value.
- Check which other gene GENEX has duplicate with this new value.
- Find if GENEX can have another value that will not cause any more duplicates. If so, go to step 7.
- If all the other values of GENEX will cause duplicates, then try another gene GENEY.
- Repeat steps 3 and 4 until exploring all the genes.
- If there is no possibility to solve the duplicates, then there is not way to solve the duplicates and we have to keep the duplicate value.
- If a value for a gene GENEM is found that will not cause more duplicates, then use this value for the gene GENEM.
- Replace the value of the gene GENE1 by the old value of the gene GENEM. This solves the duplicates.
This is an example to solve the duplicate for the solution
[3, 4, 4, 5]:
- Let's use the second gene with value 4. Because the space of this
gene is
[3, 4], then the only other value we can select is 3. - The first gene also have the value 3.
- The first gene has another value 2 that will not cause more duplicates in the solution. Then go to step 7.
- Skip.
- Skip.
- Skip.
- The value of the first gene 3 will be replaced by the new value 2. The new solution is [2, 4, 4, 5].
- Replace the value of the second gene 4 by the old value of the first gene which is 3. The new solution is [2, 3, 4, 5]. The duplicate is solved.
Example 2:
Gene space: [[0, 1],
[1, 2],
[2, 3],
[3, 4]]
Solution: [1, 2, 2, 3]The quick summary is:
- Change the value of the first gene from 1 to 0. The solution becomes [0, 2, 2, 3].
- Change the value of the second gene from 2 to 1. The solution becomes [0, 1, 2, 3]. The duplicate is solved.
Previously, the user can select the the type of the crossover, mutation,
and parent selection operators by assigning the name of the operator to
the following parameters of the pygad.GA class's constructor:
crossover_typemutation_typeparent_selection_type
This way, the user can only use the built-in functions for each of these operators.
Starting from PyGAD 2.16.0, the user can create a custom crossover, mutation, and parent selection operators and assign these functions to the above parameters. Thus, a new operator can be plugged easily into the PyGAD Lifecycle.
This is a sample code that does not use any custom function.
import pygad
import numpy
equation_inputs = [4,-2,3.5]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func)
ga_instance.run()
ga_instance.plot_fitness()This section describes the expected input parameters and outputs. For
simplicity, all of these custom functions all accept the instance of the
pygad.GA class as the last parameter.
The user-defined crossover function is a Python function that accepts 3 parameters:
- The selected parents.
- The size of the offspring as a tuple of 2 numbers: (the offspring size, number of genes).
- The instance from the
pygad.GAclass. This instance helps to retrieve any property likepopulation,gene_type,gene_space, etc.
This function should return a NumPy array of shape equal to the value passed to the second parameter.
The next code creates a template for the user-defined crossover operator. You can use any names for the parameters. Note how a NumPy array is returned.
def crossover_func(parents, offspring_size, ga_instance):
offspring = ...
...
return numpy.array(offspring)As an example, the next code creates a single-point crossover function. By randomly generating a random point (i.e. index of a gene), the function simply uses 2 parents to produce an offspring by copying the genes before the point from the first parent and the remaining from the second parent.
def crossover_func(parents, offspring_size, ga_instance):
offspring = []
idx = 0
while len(offspring) != offspring_size[0]:
parent1 = parents[idx % parents.shape[0], :].copy()
parent2 = parents[(idx + 1) % parents.shape[0], :].copy()
random_split_point = numpy.random.choice(range(offspring_size[1]))
parent1[random_split_point:] = parent2[random_split_point:]
offspring.append(parent1)
idx += 1
return numpy.array(offspring)To use this user-defined function, simply assign its name to the
crossover_type parameter in the constructor of the pygad.GA
class. The next code gives an example. In this case, the custom function
will be called in each generation rather than calling the built-in
crossover functions defined in PyGAD.
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
crossover_type=crossover_func)A user-defined mutation function/operator can be created the same way a custom crossover operator/function is created. Simply, it is a Python function that accepts 2 parameters:
- The offspring to be mutated.
- The instance from the
pygad.GAclass. This instance helps to retrieve any property likepopulation,gene_type,gene_space, etc.
The template for the user-defined mutation function is given in the next code. According to the user preference, the function should make some random changes to the genes.
def mutation_func(offspring, ga_instance):
...
return offspringThe next code builds the random mutation where a single gene from each chromosome is mutated by adding a random number between 0 and 1 to the gene's value.
def mutation_func(offspring, ga_instance):
for chromosome_idx in range(offspring.shape[0]):
random_gene_idx = numpy.random.choice(range(offspring.shape[1]))
offspring[chromosome_idx, random_gene_idx] += numpy.random.random()
return offspringHere is how this function is assigned to the mutation_type
parameter.
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
crossover_type=crossover_func,
mutation_type=mutation_func)Note that there are other things to take into consideration like:
- Making sure that each gene conforms to the data type(s) listed in the
gene_typeparameter. - If the
gene_spaceparameter is used, then the new value for the gene should conform to the values/ranges listed. - Mutating a number of genes that conforms to the parameters
mutation_percent_genes,mutation_probability, andmutation_num_genes. - Whether mutation happens with or without replacement based on the
mutation_by_replacementparameter. - The minimum and maximum values from which a random value is generated
based on the
random_mutation_min_valandrandom_mutation_max_valparameters. - Whether duplicates are allowed or not in the chromosome based on the
allow_duplicate_genesparameter.
and more.
It all depends on your objective from building the mutation function. You may neglect or consider some of the considerations according to your objective.
No much to mention about building a user-defined parent selection function as things are similar to building a crossover or mutation function. Just create a Python function that accepts 3 parameters:
- The fitness values of the current population.
- The number of parents needed.
- The instance from the
pygad.GAclass. This instance helps to retrieve any property likepopulation,gene_type,gene_space, etc.
The function should return 2 outputs:
- The selected parents as a NumPy array. Its shape is equal to (the
number of selected parents,
num_genes). Note that the number of selected parents is equal to the value assigned to the second input parameter. - The indices of the selected parents inside the population. It is a 1D list with length equal to the number of selected parents.
The outputs must be of type numpy.ndarray.
Here is a template for building a custom parent selection function.
def parent_selection_func(fitness, num_parents, ga_instance):
...
return parents, fitness_sorted[:num_parents]The next code builds the steady-state parent selection where the best
parents are selected. The number of parents is equal to the value in the
num_parents parameter.
def parent_selection_func(fitness, num_parents, ga_instance):
fitness_sorted = sorted(range(len(fitness)), key=lambda k: fitness[k])
fitness_sorted.reverse()
parents = numpy.empty((num_parents, ga_instance.population.shape[1]))
for parent_num in range(num_parents):
parents[parent_num, :] = ga_instance.population[fitness_sorted[parent_num], :].copy()
return parents, numpy.array(fitness_sorted[:num_parents])Finally, the defined function is assigned to the
parent_selection_type parameter as in the next code.
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
crossover_type=crossover_func,
mutation_type=mutation_func,
parent_selection_type=parent_selection_func)By discussing how to customize the 3 operators, the next code uses the previous 3 user-defined functions instead of the built-in functions.
import pygad
import numpy
equation_inputs = [4,-2,3.5]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
def parent_selection_func(fitness, num_parents, ga_instance):
fitness_sorted = sorted(range(len(fitness)), key=lambda k: fitness[k])
fitness_sorted.reverse()
parents = numpy.empty((num_parents, ga_instance.population.shape[1]))
for parent_num in range(num_parents):
parents[parent_num, :] = ga_instance.population[fitness_sorted[parent_num], :].copy()
return parents, numpy.array(fitness_sorted[:num_parents])
def crossover_func(parents, offspring_size, ga_instance):
offspring = []
idx = 0
while len(offspring) != offspring_size[0]:
parent1 = parents[idx % parents.shape[0], :].copy()
parent2 = parents[(idx + 1) % parents.shape[0], :].copy()
random_split_point = numpy.random.choice(range(offspring_size[1]))
parent1[random_split_point:] = parent2[random_split_point:]
offspring.append(parent1)
idx += 1
return numpy.array(offspring)
def mutation_func(offspring, ga_instance):
for chromosome_idx in range(offspring.shape[0]):
random_gene_idx = numpy.random.choice(range(offspring.shape[0]))
offspring[chromosome_idx, random_gene_idx] += numpy.random.random()
return offspring
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
crossover_type=crossover_func,
mutation_type=mutation_func,
parent_selection_type=parent_selection_func)
ga_instance.run()
ga_instance.plot_fitness()This is the same example but using methods instead of functions.
import pygad
import numpy
equation_inputs = [4,-2,3.5]
desired_output = 44
class Test:
def fitness_func(self, ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
def parent_selection_func(self, fitness, num_parents, ga_instance):
fitness_sorted = sorted(range(len(fitness)), key=lambda k: fitness[k])
fitness_sorted.reverse()
parents = numpy.empty((num_parents, ga_instance.population.shape[1]))
for parent_num in range(num_parents):
parents[parent_num, :] = ga_instance.population[fitness_sorted[parent_num], :].copy()
return parents, numpy.array(fitness_sorted[:num_parents])
def crossover_func(self, parents, offspring_size, ga_instance):
offspring = []
idx = 0
while len(offspring) != offspring_size[0]:
parent1 = parents[idx % parents.shape[0], :].copy()
parent2 = parents[(idx + 1) % parents.shape[0], :].copy()
random_split_point = numpy.random.choice(range(offspring_size[0]))
parent1[random_split_point:] = parent2[random_split_point:]
offspring.append(parent1)
idx += 1
return numpy.array(offspring)
def mutation_func(self, offspring, ga_instance):
for chromosome_idx in range(offspring.shape[0]):
random_gene_idx = numpy.random.choice(range(offspring.shape[1]))
offspring[chromosome_idx, random_gene_idx] += numpy.random.random()
return offspring
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=Test().fitness_func,
parent_selection_type=Test().parent_selection_func,
crossover_type=Test().crossover_func,
mutation_type=Test().mutation_func)
ga_instance.run()
ga_instance.plot_fitness()The gene_type parameter allows the user to control the data type for
all genes at once or each individual gene. In PyGAD
2.15.0,
the gene_type parameter also supports customizing the precision for
float data types. As a result, the gene_type parameter helps to:
- Select a data type for all genes with or without precision.
- Select a data type for each individual gene with or without precision.
Let's discuss things by examples.
The data type for all genes can be specified by assigning the numeric
data type directly to the gene_type parameter. This is an example to
make all genes of int data types.
gene_type=intGiven that the supported numeric data types of PyGAD include Python's
int and float in addition to all numeric types of NumPy,
then any of these types can be assigned to the gene_type parameter.
If no precision is specified for a float data type, then the
complete floating-point number is kept.
The next code uses an int data type for all genes where the genes in
the initial and final population are only integers.
import pygad
import numpy
equation_inputs = [4, -2, 3.5, 8, -2]
desired_output = 2671.1234
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
gene_type=int)
print("Initial Population")
print(ga_instance.initial_population)
ga_instance.run()
print("Final Population")
print(ga_instance.population)Initial Population
[[ 1 -1 2 0 -3]
[ 0 -2 0 -3 -1]
[ 0 -1 -1 2 0]
[-2 3 -2 3 3]
[ 0 0 2 -2 -2]]
Final Population
[[ 1 -1 2 2 0]
[ 1 -1 2 2 0]
[ 1 -1 2 2 0]
[ 1 -1 2 2 0]
[ 1 -1 2 2 0]]A precision can only be specified for a float data type and cannot
be specified for integers. Here is an example to use a precision of 3
for the float data type. In this case, all genes are of type
float and their maximum precision is 3.
gene_type=[float, 3]The next code uses prints the initial and final population where the
genes are of type float with precision 3.
import pygad
import numpy
equation_inputs = [4, -2, 3.5, 8, -2]
desired_output = 2671.1234
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
gene_type=[float, 3])
print("Initial Population")
print(ga_instance.initial_population)
ga_instance.run()
print("Final Population")
print(ga_instance.population)Initial Population
[[-2.417 -0.487 3.623 2.457 -2.362]
[-1.231 0.079 -1.63 1.629 -2.637]
[ 0.692 -2.098 0.705 0.914 -3.633]
[ 2.637 -1.339 -1.107 -0.781 -3.896]
[-1.495 1.378 -1.026 3.522 2.379]]
Final Population
[[ 1.714 -1.024 3.623 3.185 -2.362]
[ 0.692 -1.024 3.623 3.185 -2.362]
[ 0.692 -1.024 3.623 3.375 -2.362]
[ 0.692 -1.024 4.041 3.185 -2.362]
[ 1.714 -0.644 3.623 3.185 -2.362]]In PyGAD
2.14.0,
the gene_type parameter allows customizing the gene type for each
individual gene. This is by using a list/tuple/numpy.ndarray
with number of elements equal to the number of genes. For each element,
a type is specified for the corresponding gene.
This is an example for a 5-gene problem where different types are assigned to the genes.
gene_type=[int, float, numpy.float16, numpy.int8, float]This is a complete code that prints the initial and final population for a custom-gene data type.
import pygad
import numpy
equation_inputs = [4, -2, 3.5, 8, -2]
desired_output = 2671.1234
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
gene_type=[int, float, numpy.float16, numpy.int8, float])
print("Initial Population")
print(ga_instance.initial_population)
ga_instance.run()
print("Final Population")
print(ga_instance.population)Initial Population
[[0 0.8615522360026828 0.7021484375 -2 3.5301821368185866]
[-3 2.648189378595294 -3.830078125 1 -0.9586271572917742]
[3 3.7729827570110714 1.2529296875 -3 1.395741994211889]
[0 1.0490687178053282 1.51953125 -2 0.7243617940450235]
[0 -0.6550158436937226 -2.861328125 -2 1.8212734549263097]]
Final Population
[[3 3.7729827570110714 2.055 0 0.7243617940450235]
[3 3.7729827570110714 1.458 0 -0.14638754050305036]
[3 3.7729827570110714 1.458 0 0.0869406120516778]
[3 3.7729827570110714 1.458 0 0.7243617940450235]
[3 3.7729827570110714 1.458 0 -0.14638754050305036]]The precision can also be specified for the float data types as in
the next line where the second gene precision is 2 and last gene
precision is 1.
gene_type=[int, [float, 2], numpy.float16, numpy.int8, [float, 1]]This is a complete example where the initial and final populations are printed where the genes comply with the data types and precisions specified.
import pygad
import numpy
equation_inputs = [4, -2, 3.5, 8, -2]
desired_output = 2671.1234
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
gene_type=[int, [float, 2], numpy.float16, numpy.int8, [float, 1]])
print("Initial Population")
print(ga_instance.initial_population)
ga_instance.run()
print("Final Population")
print(ga_instance.population)Initial Population
[[-2 -1.22 1.716796875 -1 0.2]
[-1 -1.58 -3.091796875 0 -1.3]
[3 3.35 -0.107421875 1 -3.3]
[-2 -3.58 -1.779296875 0 0.6]
[2 -3.73 2.65234375 3 -0.5]]
Final Population
[[2 -4.22 3.47 3 -1.3]
[2 -3.73 3.47 3 -1.3]
[2 -4.22 3.47 2 -1.3]
[2 -4.58 3.47 3 -1.3]
[2 -3.73 3.47 3 -1.3]]This section discusses the different options to visualize the results in PyGAD through these methods:
plot_fitness()plot_genes()plot_new_solution_rate()
In the following code, the save_solutions flag is set to True
which means all solutions are saved in the solutions attribute. The
code runs for only 10 generations.
import pygad
import numpy
equation_inputs = [4, -2, 3.5, 8, -2, 3.5, 8]
desired_output = 2671.1234
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=10,
num_parents_mating=5,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
gene_space=[range(1, 10), range(10, 20), range(15, 30), range(20, 40), range(25, 50), range(10, 30), range(20, 50)],
gene_type=int,
save_solutions=True)
ga_instance.run()Let's explore how to visualize the results by the above mentioned methods.
The plot_fitness() method shows the fitness value for each
generation.
The simplest way to call this method is as follows leaving the
plot_type with its default value "plot" to create a continuous
line connecting the fitness values across all generations:
ga_instance.plot_fitness()
# ga_instance.plot_fitness(plot_type="plot")The plot_type can also be set to "scatter" to create a scatter
graph with each individual fitness represented as a dot. The size of
these dots can be changed using the linewidth parameter.
ga_instance.plot_fitness(plot_type="scatter")The third value for the plot_type parameter is "bar" to create a
bar graph with each individual fitness represented as a bar.
ga_instance.plot_fitness(plot_type="bar")The plot_new_solution_rate() method presents the number of new
solutions explored in each generation. This helps to figure out if the
genetic algorithm is able to find new solutions as an indication of more
possible evolution. If no new solutions are explored, this is an
indication that no further evolution is possible.
The plot_new_solution_rate() method accepts the same parameters as
in the plot_fitness() method with 3 possible values for
plot_type parameter.
The default value for the plot_type parameter is "plot".
ga_instance.plot_new_solution_rate()
# ga_instance.plot_new_solution_rate(plot_type="plot")The next figure shows that, for example, generation 6 has the least
number of new solutions which is 4. The number of new solutions in the
first generation is always equal to the number of solutions in the
population (i.e. the value assigned to the sol_per_pop parameter in
the constructor of the pygad.GA class) which is 10 in this example.
The previous graph can be represented as scattered points by setting
plot_type="scatter".
ga_instance.plot_new_solution_rate(plot_type="scatter")By setting plot_type="scatter", each value is represented as a
vertical bar.
ga_instance.plot_new_solution_rate(plot_type="bar")The plot_genes() method is the third option to visualize the PyGAD
results. This method has 3 control variables:
graph_type="plot": Can be"plot"(default),"boxplot", or"histogram".plot_type="plot": Identical to theplot_typeparameter explored in theplot_fitness()andplot_new_solution_rate()methods.solutions="all": Can be"all"(default) or"best".
These 3 parameters controls the style of the output figure.
The graph_type parameter selects the type of the graph which helps
to explore the gene values as:
- A normal plot.
- A histogram.
- A box and whisker plot.
The plot_type parameter works only when the type of the graph is set
to "plot".
The solutions parameter selects whether the genes come from all
solutions in the population or from just the best solutions.
When graph_type="plot", then the figure creates a normal graph where
the relationship between the gene values and the generation numbers is
represented as a continuous plot, scattered points, or bars.
Because the default value for both graph_type and plot_type is
"plot", then all of the lines below creates the same figure. This
figure is helpful to know whether a gene value lasts for more
generations as an indication of the best value for this gene. For
example, the value 16 for the gene with index 5 (at column 2 and row 2
of the next graph) lasted for 83 generations.
ga_instance.plot_genes()
ga_instance.plot_genes(graph_type="plot")
ga_instance.plot_genes(plot_type="plot")
ga_instance.plot_genes(graph_type="plot",
plot_type="plot")As the default value for the solutions parameter is "all", then
the following method calls generate the same plot.
ga_instance.plot_genes(solutions="all")
ga_instance.plot_genes(graph_type="plot",
solutions="all")
ga_instance.plot_genes(plot_type="plot",
solutions="all")
ga_instance.plot_genes(graph_type="plot",
plot_type="plot",
solutions="all")The following calls of the plot_genes() method create the same
scatter plot.
ga_instance.plot_genes(plot_type="scatter")
ga_instance.plot_genes(graph_type="plot",
plot_type="scatter",
solutions='all')ga_instance.plot_genes(plot_type="bar")
ga_instance.plot_genes(graph_type="plot",
plot_type="bar",
solutions='all')By setting graph_type to "boxplot", then a box and whisker graph
is created. Now, the plot_type parameter has no effect.
The following 2 calls of the plot_genes() method create the same
figure as the default value for the solutions parameter is
"all".
ga_instance.plot_genes(graph_type="boxplot")
ga_instance.plot_genes(graph_type="boxplot",
solutions='all')For graph_type="boxplot", then a histogram is created for each gene.
Similar to graph_type="boxplot", the plot_type parameter has no
effect.
The following 2 calls of the plot_genes() method create the same
figure as the default value for the solutions parameter is
"all".
ga_instance.plot_genes(graph_type="histogram")
ga_instance.plot_genes(graph_type="histogram",
solutions='all')All the previous figures can be created for only the best solutions by
setting solutions="best".
Starting from PyGAD 2.17.0, parallel processing becomes supported. This section explains how to use parallel processing in PyGAD.
According to the PyGAD lifecycle, parallel processing can be parallelized in only 2 operations:
- Population fitness calculation.
- Mutation.
The reason is that the calculations in these 2 operations are independent (i.e. each solution/chromosome is handled independently from the others) and can be distributed across different processes or threads.
For the mutation operation, it does not do intensive calculations on the CPU. Its calculations are simple like flipping the values of some genes from 0 to 1 or adding a random value to some genes. So, it does not take much CPU processing time. Experiments proved that parallelizing the mutation operation across the solutions increases the time instead of reducing it. This is because running multiple processes or threads adds overhead to manage them. Thus, parallel processing cannot be applied on the mutation operation.
For the population fitness calculation, parallel processing can help make a difference and reduce the processing time. But this is conditional on the type of calculations done in the fitness function. If the fitness function makes intensive calculations and takes much processing time from the CPU, then it is probably that parallel processing will help to cut down the overall time.
This section explains how parallel processing works in PyGAD and how to use parallel processing in PyGAD
Starting from PyGAD
2.17.0,
a new parameter called parallel_processing added to the constructor
of the pygad.GA class.
import pygad
...
ga_instance = pygad.GA(...,
parallel_processing=...)
...This parameter allows the user to do the following:
- Enable parallel processing.
- Select whether processes or threads are used.
- Specify the number of processes or threads to be used.
These are 3 possible values for the parallel_processing parameter:
None: (Default) It means no parallel processing is used.- A positive integer referring to the number of threads to be used (i.e. threads, not processes, are used.
list/tuple: If a list or a tuple of exactly 2 elements is assigned, then:- The first element can be either
'process'or'thread'to specify whether processes or threads are used, respectively. - The second element can be:
- A positive integer to select the maximum number of processes or threads to be used
0to indicate that 0 processes or threads are used. It means no parallel processing. This is identical to settingparallel_processing=None.Noneto use the default value as calculated by theconcurrent.futures module.
- The first element can be either
These are examples of the values assigned to the parallel_processing
parameter:
parallel_processing=4: Because the parameter is assigned a positive integer, this means parallel processing is activated where 4 threads are used.parallel_processing=["thread", 5]: Use parallel processing with 5 threads. This is identical toparallel_processing=5.parallel_processing=["process", 8]: Use parallel processing with 8 processes.parallel_processing=["process", 0]: As the second element is given the value 0, this means do not use parallel processing. This is identical toparallel_processing=None.
The examples will help you know the difference between using processes and threads. Moreover, it will give an idea when parallel processing would make a difference and reduce the time. These are dummy examples where the fitness function is made to always return 0.
The first example uses 10 genes, 5 solutions in the population where
only 3 solutions mate, and 9999 generations. The fitness function uses a
for loop with 100 iterations just to have some calculations. In the
constructor of the pygad.GA class, parallel_processing=None
means no parallel processing is used.
import pygad
import time
def fitness_func(ga_instance, solution, solution_idx):
for _ in range(99):
pass
return 0
ga_instance = pygad.GA(num_generations=9999,
num_parents_mating=3,
sol_per_pop=5,
num_genes=10,
fitness_func=fitness_func,
suppress_warnings=True,
parallel_processing=None)
if __name__ == '__main__':
t1 = time.time()
ga_instance.run()
t2 = time.time()
print("Time is", t2-t1)When parallel processing is not used, the time it takes to run the
genetic algorithm is 1.5 seconds.
In the comparison, let's do a second experiment where parallel
processing is used with 5 threads. In this case, it take 5 seconds.
...
ga_instance = pygad.GA(...,
parallel_processing=5)
...For the third experiment, processes instead of threads are used. Also,
only 99 generations are used instead of 9999. The time it takes is
99 seconds.
...
ga_instance = pygad.GA(num_generations=99,
...,
parallel_processing=["process", 5])
...This is the summary of the 3 experiments:
- No parallel processing & 9999 generations: 1.5 seconds.
- Parallel processing with 5 threads & 9999 generations: 5 seconds
- Parallel processing with 5 processes & 99 generations: 99 seconds
Because the fitness function does not need much CPU time, the normal processing takes the least time. Running processes for this simple problem takes 99 compared to only 5 seconds for threads because managing processes is much heavier than managing threads. Thus, most of the CPU time is for swapping the processes instead of executing the code.
In the second example, the loop makes 99999999 iterations and only 5 generations are used. With no parallelization, it takes 22 seconds.
import pygad
import time
def fitness_func(ga_instance, solution, solution_idx):
for _ in range(99999999):
pass
return 0
ga_instance = pygad.GA(num_generations=5,
num_parents_mating=3,
sol_per_pop=5,
num_genes=10,
fitness_func=fitness_func,
suppress_warnings=True,
parallel_processing=None)
if __name__ == '__main__':
t1 = time.time()
ga_instance.run()
t2 = time.time()
print("Time is", t2-t1)It takes 15 seconds when 10 processes are used.
...
ga_instance = pygad.GA(...,
parallel_processing=["process", 10])
...This is compared to 20 seconds when 10 threads are used.
...
ga_instance = pygad.GA(...,
parallel_processing=["thread", 10])
...Based on the second example, using parallel processing with 10 processes takes the least time because there is much CPU work done. Generally, processes are preferred over threads when most of the work in on the CPU. Threads are preferred over processes in some situations like doing input/output operations.
Before releasing PyGAD 2.17.0, László Fazekas wrote an article to parallelize the fitness function with PyGAD. Check it: How Genetic Algorithms Can Compete with Gradient Descent and Backprop.
In PyGAD
2.19.0,
a new method called summary() is supported. It prints a Keras-like
summary of the PyGAD lifecycle showing the steps, callback functions,
parameters, etc.
This method accepts the following parameters:
line_length=70: An integer representing the length of the single line in characters.fill_character=" ": A character to fill the lines.line_character="-": A character for creating a line separator.line_character2="=": A secondary character to create a line separator.columns_equal_len=False: The table rows are split into equal-sized columns or split subjective to the width needed.print_step_parameters=True: Whether to print extra parameters about each step inside the step. Ifprint_step_parameters=Falseandprint_parameters_summary=True, then the parameters of each step are printed at the end of the table.print_parameters_summary=True: Whether to print parameters summary at the end of the table. Ifprint_step_parameters=False, then the parameters of each step are printed at the end of the table too.
This is a quick example to create a PyGAD example.
import pygad
import numpy
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
def genetic_fitness(solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
def on_gen(ga):
pass
def on_crossover_callback(a, b):
pass
ga_instance = pygad.GA(num_generations=100,
num_parents_mating=10,
sol_per_pop=20,
num_genes=len(function_inputs),
on_crossover=on_crossover_callback,
on_generation=on_gen,
parallel_processing=2,
stop_criteria="reach_10",
fitness_batch_size=4,
crossover_probability=0.4,
fitness_func=genetic_fitness)Then call the summary() method to print the summary with the default
parameters. Note that entries for the crossover and generation callback
function are created because their callback functions are implemented
through the on_crossover_callback() and on_gen(), respectively.
ga_instance.summary()----------------------------------------------------------------------
PyGAD Lifecycle
======================================================================
Step Handler Output Shape
======================================================================
Fitness Function genetic_fitness() (1)
Fitness batch size: 4
----------------------------------------------------------------------
Parent Selection steady_state_selection() (10, 6)
Number of Parents: 10
----------------------------------------------------------------------
Crossover single_point_crossover() (10, 6)
Crossover probability: 0.4
----------------------------------------------------------------------
On Crossover on_crossover_callback() None
----------------------------------------------------------------------
Mutation random_mutation() (10, 6)
Mutation Genes: 1
Random Mutation Range: (-1.0, 1.0)
Mutation by Replacement: False
Allow Duplicated Genes: True
----------------------------------------------------------------------
On Generation on_gen() None
Stop Criteria: [['reach', 10.0]]
----------------------------------------------------------------------
======================================================================
Population Size: (20, 6)
Number of Generations: 100
Initial Population Range: (-4, 4)
Keep Elitism: 1
Gene DType: [<class 'float'>, None]
Parallel Processing: ['thread', 2]
Save Best Solutions: False
Save Solutions: False
======================================================================We can set the print_step_parameters and
print_parameters_summary parameters to False to not print the
parameters.
ga_instance.summary(print_step_parameters=False,
print_parameters_summary=False)----------------------------------------------------------------------
PyGAD Lifecycle
======================================================================
Step Handler Output Shape
======================================================================
Fitness Function genetic_fitness() (1)
----------------------------------------------------------------------
Parent Selection steady_state_selection() (10, 6)
----------------------------------------------------------------------
Crossover single_point_crossover() (10, 6)
----------------------------------------------------------------------
On Crossover on_crossover_callback() None
----------------------------------------------------------------------
Mutation random_mutation() (10, 6)
----------------------------------------------------------------------
On Generation on_gen() None
----------------------------------------------------------------------
======================================================================In PyGAD
3.0.0,
the print() statement is no longer used and the outputs are printed
using the logging
module. A a new parameter called logger is supported to accept the
user-defined logger.
import logging
logger = ...
ga_instance = pygad.GA(...,
logger=logger,
...)The default value for this parameter is None. If there is no logger
passed (i.e. logger=None), then a default logger is created to log
the messages to the console exactly like how the print() statement
works.
Some advantages of using the the
logging module
instead of the print() statement are:
- The user has more control over the printed messages specially if there is a project that uses multiple modules where each module prints its messages. A logger can organize the outputs.
- Using the proper
Handler, the user can log the output messages to files and not only restricted to printing it to the console. So, it is much easier to record the outputs. - The format of the printed messages can be changed by customizing the
Formatterassigned to the Logger.
This section gives some quick examples to use the logging module and
then gives an example to use the logger with PyGAD.
This is an example to create a logger to log the messages to the console.
import logging
# Create a logger
logger = logging.getLogger(__name__)
# Set the logger level to debug so that all the messages are printed.
logger.setLevel(logging.DEBUG)
# Create a stream handler to log the messages to the console.
stream_handler = logging.StreamHandler()
# Set the handler level to debug.
stream_handler.setLevel(logging.DEBUG)
# Create a formatter
formatter = logging.Formatter('%(message)s')
# Add the formatter to handler.
stream_handler.setFormatter(formatter)
# Add the stream handler to the logger
logger.addHandler(stream_handler)Now, we can log messages to the console with the format specified in the
Formatter.
logger.debug('Debug message.')
logger.info('Info message.')
logger.warning('Warn message.')
logger.error('Error message.')
logger.critical('Critical message.')The outputs are identical to those returned using the print()
statement.
Debug message. Info message. Warn message. Error message. Critical message.
By changing the format of the output messages, we can have more information about each message.
formatter = logging.Formatter('%(asctime)s %(levelname)s: %(message)s', datefmt='%Y-%m-%d %H:%M:%S')This is a sample output.
2023-04-03 18:46:27 DEBUG: Debug message.
2023-04-03 18:46:27 INFO: Info message.
2023-04-03 18:46:27 WARNING: Warn message.
2023-04-03 18:46:27 ERROR: Error message.
2023-04-03 18:46:27 CRITICAL: Critical message.Note that you may need to clear the handlers after finishing the execution. This is to make sure no cached handlers are used in the next run. If the cached handlers are not cleared, then the single output message may be repeated.
logger.handlers.clear()This is another example to log the messages to a file named
logfile.txt. The formatter prints the following about each message:
- The date and time at which the message is logged.
- The log level.
- The message.
- The path of the file.
- The lone number of the log message.
import logging
level = logging.DEBUG
name = 'logfile.txt'
logger = logging.getLogger(name)
logger.setLevel(level)
file_handler = logging.FileHandler(name, 'a+', 'utf-8')
file_handler.setLevel(logging.DEBUG)
file_format = logging.Formatter('%(asctime)s %(levelname)s: %(message)s - %(pathname)s:%(lineno)d', datefmt='%Y-%m-%d %H:%M:%S')
file_handler.setFormatter(file_format)
logger.addHandler(file_handler)This is how the outputs look like.
2023-04-03 18:54:03 DEBUG: Debug message. - c:\users\agad069\desktop\logger\example2.py:46
2023-04-03 18:54:03 INFO: Info message. - c:\users\agad069\desktop\logger\example2.py:47
2023-04-03 18:54:03 WARNING: Warn message. - c:\users\agad069\desktop\logger\example2.py:48
2023-04-03 18:54:03 ERROR: Error message. - c:\users\agad069\desktop\logger\example2.py:49
2023-04-03 18:54:03 CRITICAL: Critical message. - c:\users\agad069\desktop\logger\example2.py:50Consider clearing the handlers if necessary.
logger.handlers.clear()This is an example to create a single Logger associated with 2 handlers:
- A file handler.
- A stream handler.
import logging
level = logging.DEBUG
name = 'logfile.txt'
logger = logging.getLogger(name)
logger.setLevel(level)
file_handler = logging.FileHandler(name,'a+','utf-8')
file_handler.setLevel(logging.DEBUG)
file_format = logging.Formatter('%(asctime)s %(levelname)s: %(message)s - %(pathname)s:%(lineno)d', datefmt='%Y-%m-%d %H:%M:%S')
file_handler.setFormatter(file_format)
logger.addHandler(file_handler)
console_handler = logging.StreamHandler()
console_handler.setLevel(logging.INFO)
console_format = logging.Formatter('%(message)s')
console_handler.setFormatter(console_format)
logger.addHandler(console_handler)When a log message is executed, then it is both printed to the console
and saved in the logfile.txt.
Consider clearing the handlers if necessary.
logger.handlers.clear()To use the logger in PyGAD, just create your custom logger and pass it
to the logger parameter.
import logging
import pygad
import numpy
level = logging.DEBUG
name = 'logfile.txt'
logger = logging.getLogger(name)
logger.setLevel(level)
file_handler = logging.FileHandler(name,'a+','utf-8')
file_handler.setLevel(logging.DEBUG)
file_format = logging.Formatter('%(asctime)s %(levelname)s: %(message)s', datefmt='%Y-%m-%d %H:%M:%S')
file_handler.setFormatter(file_format)
logger.addHandler(file_handler)
console_handler = logging.StreamHandler()
console_handler.setLevel(logging.INFO)
console_format = logging.Formatter('%(message)s')
console_handler.setFormatter(console_format)
logger.addHandler(console_handler)
equation_inputs = [4, -2, 8]
desired_output = 2671.1234
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
def on_generation(ga_instance):
ga_instance.logger.info("Generation = {generation}".format(generation=ga_instance.generations_completed))
ga_instance.logger.info("Fitness = {fitness}".format(fitness=ga_instance.best_solution(pop_fitness=ga_instance.last_generation_fitness)[1]))
ga_instance = pygad.GA(num_generations=10,
sol_per_pop=40,
num_parents_mating=2,
keep_parents=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
on_generation=on_generation,
logger=logger)
ga_instance.run()
logger.handlers.clear()By executing this code, the logged messages are printed to the console and also saved in the text file.
2023-04-03 19:04:27 INFO: Generation = 1
2023-04-03 19:04:27 INFO: Fitness = 0.00038086960368076276
2023-04-03 19:04:27 INFO: Generation = 2
2023-04-03 19:04:27 INFO: Fitness = 0.00038214871408010853
2023-04-03 19:04:27 INFO: Generation = 3
2023-04-03 19:04:27 INFO: Fitness = 0.0003832795907974678
2023-04-03 19:04:27 INFO: Generation = 4
2023-04-03 19:04:27 INFO: Fitness = 0.00038398612055017196
2023-04-03 19:04:27 INFO: Generation = 5
2023-04-03 19:04:27 INFO: Fitness = 0.00038442348890867516
2023-04-03 19:04:27 INFO: Generation = 6
2023-04-03 19:04:27 INFO: Fitness = 0.0003854406039137763
2023-04-03 19:04:27 INFO: Generation = 7
2023-04-03 19:04:27 INFO: Fitness = 0.00038646083174063284
2023-04-03 19:04:27 INFO: Generation = 8
2023-04-03 19:04:27 INFO: Fitness = 0.0003875169193024936
2023-04-03 19:04:27 INFO: Generation = 9
2023-04-03 19:04:27 INFO: Fitness = 0.0003888816727311021
2023-04-03 19:04:27 INFO: Generation = 10
2023-04-03 19:04:27 INFO: Fitness = 0.000389832593101348PyGAD can be used to solve both deterministic and non-deterministic problems. Deterministic are those that return the same fitness for the same solution. For non-deterministic problems, a different fitness value would be returned for the same solution.
By default, PyGAD settings are set to solve deterministic problems. PyGAD can save the explored solutions and their fitness to reuse in the future. These instances attributes can save the solutions:
solutions: Exists ifsave_solutions=True.best_solutions: Exists ifsave_best_solutions=True.last_generation_elitism: Exists ifkeep_elitism> 0.last_generation_parents: Exists ifkeep_parents> 0 orkeep_parents=-1.
To configure PyGAD for non-deterministic problems, we have to disable saving the previous solutions. This is by setting these parameters:
keep_elisitm=0keep_parents=0keep_solutions=Falsekeep_best_solutions=False
import pygad
...
ga_instance = pygad.GA(...,
keep_elitism=0,
keep_parents=0,
save_solutions=False,
save_best_solutions=False,
...)This way PyGAD will not save any explored solution and thus the fitness function have to be called for each individual solution.
It may happen that a previously explored solution in generation X is explored again in another generation Y (where Y > X). For some problems, calling the fitness function takes much time.
For deterministic problems, it is better to not call the fitness function for an already explored solutions. Instead, reuse the fitness of the old solution. PyGAD supports some options to help you save time calling the fitness function for a previously explored solution.
The parameters explored in this section can be set in the constructor of
the pygad.GA class.
The cal_pop_fitness() method of the pygad.GA class checks these
parameters to see if there is a possibility of reusing the fitness
instead of calling the fitness function.
It defaults to False. If set to True, then the population of
each generation is saved into the solutions attribute of the
pygad.GA instance. In other words, every single solution is saved in
the solutions attribute.
It defaults to False. If True, then it only saves the best
solution in every generation.
It accepts an integer and defaults to 1. If set to a positive integer, then it keeps the elitism of one generation available in the next generation.
It accepts an integer and defaults to -1. It set to -1 or a positive
integer, then it keeps the parents of one generation available in the
next generation.
PyGAD has a parameter called keep_elitism which defaults to 1. This
parameter defines the number of best solutions in generation X to
keep in the next generation X+1. The best solutions are just copied
from generation X to generation X+1 without making any change.
ga_instance = pygad.GA(...,
keep_elitism=1,
...)The best solutions are copied at the beginning of the population. If
keep_elitism=1, this means the best solution in generation X is kept
in the next generation X+1 at index 0 of the population. If
keep_elitism=2, this means the 2 best solutions in generation X are
kept in the next generation X+1 at indices 0 and 1 of the population of
generation 1.
Because the fitness of these best solutions are already calculated in generation X, then their fitness values will not be recalculated at generation X+1 (i.e. the fitness function will not be called for these solutions again). Instead, their fitness values are just reused. This is why you see that no solution with index 0 is passed to the fitness function.
To force calling the fitness function for each solution in every
generation, consider setting keep_elitism and keep_parents to 0.
Moreover, keep the 2 parameters save_solutions and
save_best_solutions to their default value False.
ga_instance = pygad.GA(...,
keep_elitism=0,
keep_parents=0,
save_solutions=False,
save_best_solutions=False,
...)In PyGAD
2.19.0,
a new optional parameter called fitness_batch_size is supported. A
new optional parameter called fitness_batch_size is supported to
calculate the fitness function in batches. Thanks to Linan
Qiu for opening the GitHub issue
#136.
Its values can be:
1orNone: If thefitness_batch_sizeparameter is assigned the value1orNone(default), then the normal flow is used where the fitness function is called for each individual solution. That is if there are 15 solutions, then the fitness function is called 15 times.1 < fitness_batch_size <= sol_per_pop: If thefitness_batch_sizeparameter is assigned a value satisfying this condition1 < fitness_batch_size <= sol_per_pop, then the solutions are grouped into batches of sizefitness_batch_sizeand the fitness function is called once for each batch. In this case, the fitness function must return a list/tuple/numpy.ndarray with a length equal to the number of solutions passed.
This is an example where the fitness_batch_size parameter is given
the value None (which is the default value). This is equivalent to
using the value 1. In this case, the fitness function will be called
for each solution. This means the fitness function fitness_func will
receive only a single solution. This is an example of the passed
arguments to the fitness function:
solution: [ 2.52860734, -0.94178795, 2.97545704, 0.84131987, -3.78447118, 2.41008358] solution_idx: 3
The fitness function also must return a single numeric value as the fitness for the passed solution.
As we have a population of 20 solutions, then the fitness function
is called 20 times per generation. For 5 generations, then the fitness
function is called 20*5 = 100 times. In PyGAD, the fitness function
is called after the last generation too and this adds additional 20
times. So, the total number of calls to the fitness function is
20*5 + 20 = 120.
Note that the keep_elitism and keep_parents parameters are set
to 0 to make sure no fitness values are reused and to force calling
the fitness function for each individual solution.
import pygad
import numpy
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
number_of_calls = 0
def fitness_func(ga_instance, solution, solution_idx):
global number_of_calls
number_of_calls = number_of_calls + 1
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
ga_instance = pygad.GA(num_generations=5,
num_parents_mating=10,
sol_per_pop=20,
fitness_func=fitness_func,
fitness_batch_size=None,
# fitness_batch_size=1,
num_genes=len(function_inputs),
keep_elitism=0,
keep_parents=0)
ga_instance.run()
print(number_of_calls)120
This is an example where the fitness_batch_size parameter is used
and assigned the value 4. This means the solutions will be grouped
into batches of 4 solutions. The fitness function will be called
once for each patch (i.e. called once for each 4 solutions).
This is an example of the arguments passed to it:
solutions:
[[ 3.1129432 -0.69123589 1.93792414 2.23772968 -1.54616001 -0.53930799]
[ 3.38508121 0.19890812 1.93792414 2.23095014 -3.08955597 3.10194128]
[ 2.37079504 -0.88819803 2.97545704 1.41742256 -3.95594055 2.45028256]
[ 2.52860734 -0.94178795 2.97545704 0.84131987 -3.78447118 2.41008358]]
solutions_indices:
[16, 17, 18, 19]As we have 20 solutions, then there are 20/4 = 5 patches. As a
result, the fitness function is called only 5 times per generation
instead of 20. For each call to the fitness function, it receives a
batch of 4 solutions.
As we have 5 generations, then the function will be called 5*5 = 25
times. Given the call to the fitness function after the last generation,
then the total number of calls is 5*5 + 5 = 30.
import pygad
import numpy
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
number_of_calls = 0
def fitness_func_batch(ga_instance, solutions, solutions_indices):
global number_of_calls
number_of_calls = number_of_calls + 1
batch_fitness = []
for solution in solutions:
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
batch_fitness.append(fitness)
return batch_fitness
ga_instance = pygad.GA(num_generations=5,
num_parents_mating=10,
sol_per_pop=20,
fitness_func=fitness_func_batch,
fitness_batch_size=4,
num_genes=len(function_inputs),
keep_elitism=0,
keep_parents=0)
ga_instance.run()
print(number_of_calls)30
When batch fitness calculation is used, then we saved 120 - 30 = 90
calls to the fitness function.
In PyGAD 2.19.0, it is possible to pass user-defined functions or methods to the following parameters:
fitness_funcon_starton_fitnesson_parentson_crossoveron_mutationon_generationon_stop
This section gives 2 examples to assign these parameters user-defined:
- Functions.
- Methods.
This is a dummy example where the fitness function returns a random
value. Note that the instance of the pygad.GA class is passed as the
last parameter of all functions.
import pygad
import numpy
def fitness_func(ga_instanse, solution, solution_idx):
return numpy.random.rand()
def on_start(ga_instanse):
print("on_start")
def on_fitness(ga_instanse, last_gen_fitness):
print("on_fitness")
def on_parents(ga_instanse, last_gen_parents):
print("on_parents")
def on_crossover(ga_instanse, last_gen_offspring):
print("on_crossover")
def on_mutation(ga_instanse, last_gen_offspring):
print("on_mutation")
def on_generation(ga_instanse):
print("on_generation\n")
def on_stop(ga_instanse, last_gen_fitness):
print("on_stop")
ga_instance = pygad.GA(num_generations=5,
num_parents_mating=4,
sol_per_pop=10,
num_genes=2,
on_start=on_start,
on_fitness=on_fitness,
on_parents=on_parents,
on_crossover=on_crossover,
on_mutation=on_mutation,
on_generation=on_generation,
on_stop=on_stop,
fitness_func=fitness_func)
ga_instance.run()The next example has all the method defined inside the class Test.
All of the methods accept an additional parameter representing the
method's object of the class Test.
All methods accept self as the first parameter and the instance of
the pygad.GA class as the last parameter.
import pygad
import numpy
class Test:
def fitness_func(self, ga_instanse, solution, solution_idx):
return numpy.random.rand()
def on_start(self, ga_instanse):
print("on_start")
def on_fitness(self, ga_instanse, last_gen_fitness):
print("on_fitness")
def on_parents(self, ga_instanse, last_gen_parents):
print("on_parents")
def on_crossover(self, ga_instanse, last_gen_offspring):
print("on_crossover")
def on_mutation(self, ga_instanse, last_gen_offspring):
print("on_mutation")
def on_generation(self, ga_instanse):
print("on_generation\n")
def on_stop(self, ga_instanse, last_gen_fitness):
print("on_stop")
ga_instance = pygad.GA(num_generations=5,
num_parents_mating=4,
sol_per_pop=10,
num_genes=2,
on_start=Test().on_start,
on_fitness=Test().on_fitness,
on_parents=Test().on_parents,
on_crossover=Test().on_crossover,
on_mutation=Test().on_mutation,
on_generation=Test().on_generation,
on_stop=Test().on_stop,
fitness_func=Test().fitness_func)
ga_instance.run()This section gives the complete code of some examples that use
pygad. Each subsection builds a different example.
This example is discussed in the Steps to Use PyGAD section which optimizes a linear model. Its complete code is listed below.
import pygad
import numpy
"""
Given the following function:
y = f(w1:w6) = w1x1 + w2x2 + w3x3 + w4x4 + w5x5 + 6wx6
where (x1,x2,x3,x4,x5,x6)=(4,-2,3.5,5,-11,-4.7) and y=44
What are the best values for the 6 weights (w1 to w6)? We are going to use the genetic algorithm to optimize this function.
"""
function_inputs = [4,-2,3.5,5,-11,-4.7] # Function inputs.
desired_output = 44 # Function output.
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
num_generations = 100 # Number of generations.
num_parents_mating = 10 # Number of solutions to be selected as parents in the mating pool.
sol_per_pop = 20 # Number of solutions in the population.
num_genes = len(function_inputs)
last_fitness = 0
def on_generation(ga_instance):
global last_fitness
print("Generation = {generation}".format(generation=ga_instance.generations_completed))
print("Fitness = {fitness}".format(fitness=ga_instance.best_solution(pop_fitness=ga_instance.last_generation_fitness)[1]))
print("Change = {change}".format(change=ga_instance.best_solution(pop_fitness=ga_instance.last_generation_fitness)[1] - last_fitness))
last_fitness = ga_instance.best_solution(pop_fitness=ga_instance.last_generation_fitness)[1]
ga_instance = pygad.GA(num_generations=num_generations,
num_parents_mating=num_parents_mating,
sol_per_pop=sol_per_pop,
num_genes=num_genes,
fitness_func=fitness_func,
on_generation=on_generation)
# Running the GA to optimize the parameters of the function.
ga_instance.run()
ga_instance.plot_fitness()
# Returning the details of the best solution.
solution, solution_fitness, solution_idx = ga_instance.best_solution(ga_instance.last_generation_fitness)
print("Parameters of the best solution : {solution}".format(solution=solution))
print("Fitness value of the best solution = {solution_fitness}".format(solution_fitness=solution_fitness))
print("Index of the best solution : {solution_idx}".format(solution_idx=solution_idx))
prediction = numpy.sum(numpy.array(function_inputs)*solution)
print("Predicted output based on the best solution : {prediction}".format(prediction=prediction))
if ga_instance.best_solution_generation != -1:
print("Best fitness value reached after {best_solution_generation} generations.".format(best_solution_generation=ga_instance.best_solution_generation))
# Saving the GA instance.
filename = 'genetic' # The filename to which the instance is saved. The name is without extension.
ga_instance.save(filename=filename)
# Loading the saved GA instance.
loaded_ga_instance = pygad.load(filename=filename)
loaded_ga_instance.plot_fitness()This project reproduces a single image using PyGAD by evolving pixel values. This project works with both color and gray images. Check this project at GitHub: https://github.com/ahmedfgad/GARI.
For more information about this project, read this tutorial titled Reproducing Images using a Genetic Algorithm with Python available at these links:
- Heartbeat: https://heartbeat.fritz.ai/reproducing-images-using-a-genetic-algorithm-with-python-91fc701ff84
- LinkedIn: https://www.linkedin.com/pulse/reproducing-images-using-genetic-algorithm-python-ahmed-gad
The steps to follow in order to reproduce an image are as follows:
- Read an image
- Prepare the fitness function
- Create an instance of the pygad.GA class with the appropriate parameters
- Run PyGAD
- Plot results
- Calculate some statistics
The next sections discusses the code of each of these steps.
There is an image named fruit.jpg in the GARI
project which is read according
to the next code.
import imageio
import numpy
target_im = imageio.imread('fruit.jpg')
target_im = numpy.asarray(target_im/255, dtype=float)Here is the read image.
Based on the chromosome representation used in the example, the pixel values can be either in the 0-255, 0-1, or any other ranges.
Note that the range of pixel values affect other parameters like the range from which the random values are selected during mutation and also the range of the values used in the initial population. So, be consistent.
The next code creates a function that will be used as a fitness function
for calculating the fitness value for each solution in the population.
This function must be a maximization function that accepts 3 parameters
representing the instance of the pygad.GA class, a solution, and its
index. It returns a value representing the fitness value.
import gari
target_chromosome = gari.img2chromosome(target_im)
def fitness_fun(ga_instance, solution, solution_idx):
fitness = numpy.sum(numpy.abs(target_chromosome-solution))
# Negating the fitness value to make it increasing rather than decreasing.
fitness = numpy.sum(target_chromosome) - fitness
return fitnessThe fitness value is calculated using the sum of absolute difference
between genes values in the original and reproduced chromosomes. The
gari.img2chromosome() function is called before the fitness function
to represent the image as a vector because the genetic algorithm can
work with 1D chromosomes.
The implementation of the gari module is available at the GARI
GitHub
project and
its code is listed below.
import numpy
import functools
import operator
def img2chromosome(img_arr):
return numpy.reshape(a=img_arr, newshape=(functools.reduce(operator.mul, img_arr.shape)))
def chromosome2img(vector, shape):
if len(vector) != functools.reduce(operator.mul, shape):
raise ValueError("A vector of length {vector_length} into an array of shape {shape}.".format(vector_length=len(vector), shape=shape))
return numpy.reshape(a=vector, newshape=shape)It is very important to use random mutation and set the
mutation_by_replacement to True. Based on the range of pixel
values, the values assigned to the init_range_low,
init_range_high, random_mutation_min_val, and
random_mutation_max_val parameters should be changed.
If the image pixel values range from 0 to 255, then set
init_range_low and random_mutation_min_val to 0 as they are but
change init_range_high and random_mutation_max_val to 255.
Feel free to change the other parameters or add other parameters. Please check the PyGAD's documentation for the full list of parameters.
import pygad
ga_instance = pygad.GA(num_generations=20000,
num_parents_mating=10,
fitness_func=fitness_fun,
sol_per_pop=20,
num_genes=target_im.size,
init_range_low=0.0,
init_range_high=1.0,
mutation_percent_genes=0.01,
mutation_type="random",
mutation_by_replacement=True,
random_mutation_min_val=0.0,
random_mutation_max_val=1.0)Simply, call the run() method to run PyGAD.
ga_instance.run()After the run() method completes, the fitness values of all
generations can be viewed in a plot using the plot_fitness() method.
ga_instance.plot_fitness()Here is the plot after 20,000 generations.
Here is some information about the best solution.
# Returning the details of the best solution.
solution, solution_fitness, solution_idx = ga_instance.best_solution()
print("Fitness value of the best solution = {solution_fitness}".format(solution_fitness=solution_fitness))
print("Index of the best solution : {solution_idx}".format(solution_idx=solution_idx))
if ga_instance.best_solution_generation != -1:
print("Best fitness value reached after {best_solution_generation} generations.".format(best_solution_generation=ga_instance.best_solution_generation))
result = gari.chromosome2img(solution, target_im.shape)
matplotlib.pyplot.imshow(result)
matplotlib.pyplot.title("PyGAD & GARI for Reproducing Images")
matplotlib.pyplot.show()The solution reached after the 20,000 generations is shown below.
After more generations, the result can be enhanced like what shown below.
The results can also be enhanced by changing the parameters passed to
the constructor of the pygad.GA class.
Here is how the image is evolved from generation 0 to generation 20,000s.
Generation 0
Generation 1,000
Generation 2,500
Generation 4,500
Generation 7,000
Generation 8,000
Generation 20,000
For a 2-cluster problem, the code is available here. For a 3-cluster problem, the code is here. The 2 examples are using artificial samples.
Soon a tutorial will be published at Paperspace to explain how clustering works using the genetic algorithm with examples in PyGAD.
The code is available the CoinTex GitHub project. CoinTex is an Android game written in Python using the Kivy framework. Find CoinTex at Google Play: https://play.google.com/store/apps/details?id=coin.tex.cointexreactfast
Check this Paperspace tutorial for how the genetic algorithm plays CoinTex: https://blog.paperspace.com/building-agent-for-cointex-using-genetic-algorithm. Check also this YouTube video showing the genetic algorithm while playing CoinTex.